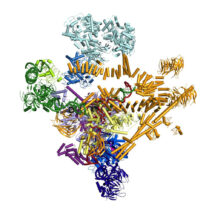
 The spliceosome is a molecular machine, which together with RNA polymerases and ribosomes plays a critical role in basic gene expression. Due to its highly dynamic nature the structure of the spliceosome has remained elusive until now. Research by Kiyoshi Nagai’s group, in the LMB’s Structural Studies Division, has for the first time captured the spliceosome in a fully active, substrate-bound state, immediately after first catalytic reaction. This work represents the first step forward to see how this machine is made and how it functions.
The spliceosome is a molecular machine, which together with RNA polymerases and ribosomes plays a critical role in basic gene expression. Due to its highly dynamic nature the structure of the spliceosome has remained elusive until now. Research by Kiyoshi Nagai’s group, in the LMB’s Structural Studies Division, has for the first time captured the spliceosome in a fully active, substrate-bound state, immediately after first catalytic reaction. This work represents the first step forward to see how this machine is made and how it functions.
The majority of eukaryotic genes contain non-coding regions called introns, which are transcribed into messenger RNA precursors (pre-mRNA). Introns must be removed from pre-mRNA to form continuous protein coding sequences before mRNAs are used for protein synthesis. This intron removal process is known as pre-mRNA splicing. Pre-mRNA splicing is catalysed by the spliceosome and proceeds by a two-step trans-esterification mechanism, analogous to group II intron self-splicing. The spliceosome is assembled on pre-mRNA by the ordered addition of small nuclear ribonucleoprotein particles (snRNPs) and numerous proteins, followed by several remodelling events before it achieves catalytic competence.
Wojciech Galej, Max Wilkinson and Sebastian Fica in Kiyoshi’s group with the help of Chris Oubridge and Andy Newman, have solved a high resolution cryo-EM structure of the spliceosome captured immediately after the first step of catalysis. They have built an atomic model for this nearly 2MDa complex composed of 44 subunits including proteins and RNAs. Most importantly, the data allowed them to visualise for the very first time the pre-mRNA substrate inside the spliceosome and its interactions with the RNA catalytic core and surrounding proteins. This has provided important mechanistic insights into RNA-based splicing catalysis. It has consolidated nearly three decades of biochemistry and genetics on the spliceosome and indicates new research directions to be explored.
Malfunction of the spliceosome is implicated in several diseases including degenerative eye disease, retinitis pigmentosa, and some cancers. In future, it may be possible to understand how mutations in this machine can lead to diseases and offer therapeutic treatments to restore the correct functioning of the spliceosome.
This work was funded by the MRC and, Sebstian was supported by EMBO and EU Marie Sklodowska-Curie fellowships and Max a Rutherford Memorial Cambridge Scholarship.
Further references:
Kiyoshi’s group page
Paper in Nature
Nature News & Views
Previous Insight on Research article: Cryo-EM sheds new light on the spliceosome activation process.