The Carter lab studies dyneins, a family of microtubule motor proteins. We focus primarily on cytoplasmic dynein, the motor responsible for much of the long-range, minus-end-directed transport within cells. This transport system is frequently hijacked by viruses, and defects in dynein function are linked to neurodegenerative and neurodevelopmental disorders.
We combine structural biology (cryo-EM and cryo-ET), in vitro reconstitution, single-molecule microscopy and complementary cell biological approaches to understand how dynein is regulated, how it recognises and transports such a diverse range of cargos, and how its activity is coordinated with oppositely directed kinesin motors.
Past highlights include: X-ray crystal structures of the dynein motor domain that explain how it couples ATP hydrolysis to movement (Schmidt et al 2015); the discovery that coiled coil cargo adaptors activate dynein’s long-distance movement (Schlager et al 2014); cryoEM structures of dynein and its co-factor dynactin which reveal how dynein is activated (Urnavicius et al 2015, Zhang et al 2018).
Current Projects
A key feature of cytoplasmic dynein is the large number of roles it plays in the cell. It transports many different membrane organelles and everything from aggregated proteins to viruses. It also has critical roles in cell division. Our current focus is to work out what components are required to recruit dynein to specific cargos and how the transport is controlled. Our approach involves structure determination, cell biology approaches, in vitro reconstitution of dynein/cargo complexes, as well as direct visualization of cargos inside the axons of neurons by cryo-electron tomography.
Recent Research Highlights
Dynein-dynactin on microtubules
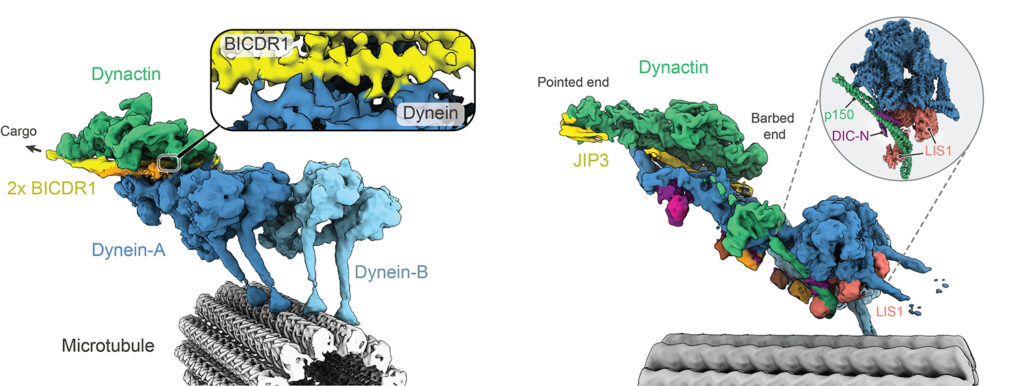
Our early cryo-EM structures showed how the tails of dynein bind to dynactin and a coiled coil adaptor. To improve the resolution, we established how to solve the whole dynein-dynactin-adaptor complex bound to microtubules. A key technical advance was subtracting the signal of the microtubule from the raw images.
Our first structure showed the unanticipated presence of two copies of the adaptor BICDR1 and allowed us to identify the full adaptor motif required to bind dynein (Chaaban and Carter, 2022).
We next determined how the adaptor JIP3 specifies dynein–dynactin binding despite being only half the length of many other adaptors. This work enabled our collaborator Reto Gassmann (i3S, Portugal) to identify two novel “short coiled-coil” adaptors, CDR1 and CDR2, which bind the endoplasmic reticulum (Teixeira et al, 2025).
The JIP3-bound structure was solved in the presence of the dynein regulator LIS1, mutations in which lead to the neurodevelopmental disorder Lissencephaly. To our surprise LIS1 contacted the long p150 arm of dynactin, binding it along the length of the dynein. We proposed a model for how this accounts for LIS1’s ability to stimulate formation of dynein-dynactin complexes (Singh et al, 2024).
Dynein and Kinesins
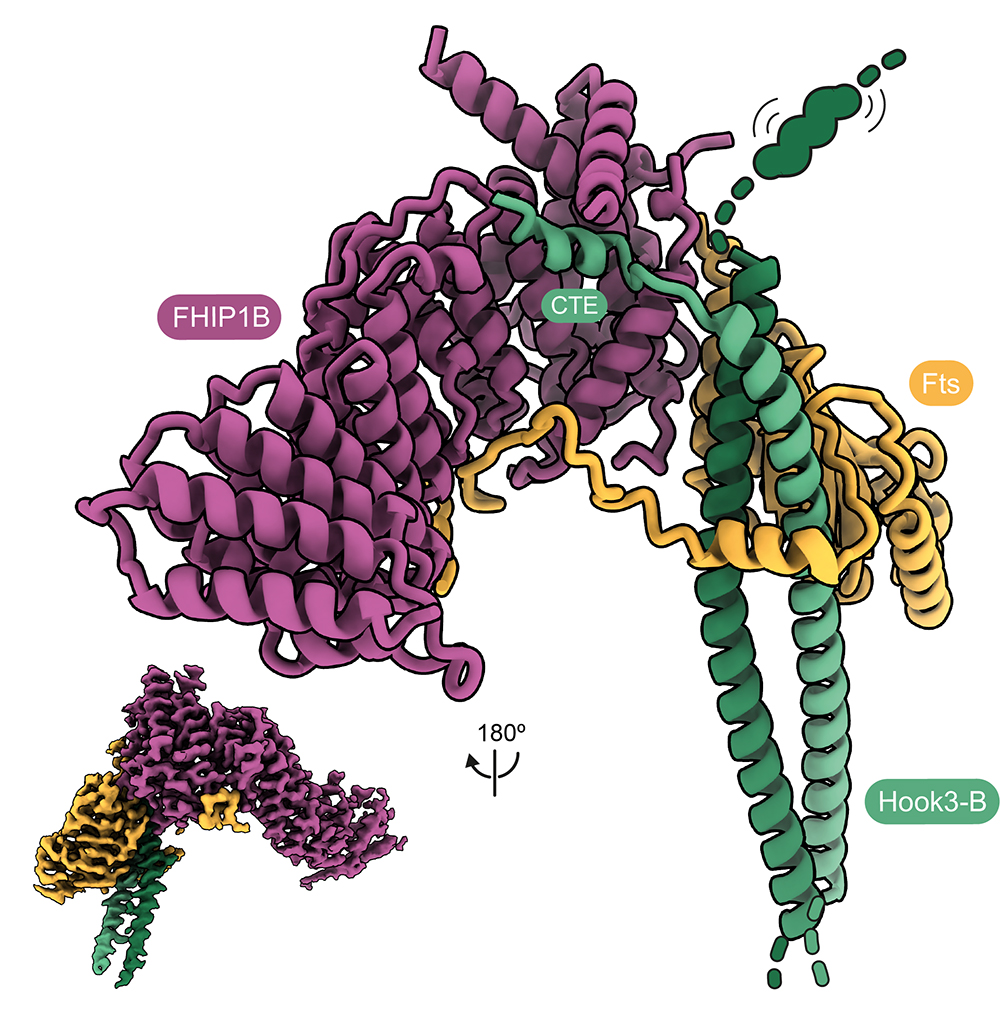
Cargo transport in cells frequently requires coordination between dynein and the opposite direction kinesin motors. Many dynein adaptors are also known to bind kinesins. We collaborated with Anne Straube (Warwick) to show how the adaptor HOOK3 is activated for binding dynein-dynactin by its interaction with the kinesin-3 KIF1C. This provided a molecular explanation for why cargo movements can co-depend on the presence of both types of motors. We further showed how HOOK3 binds two proteins (FTS and FHIP) which tether it to cargos, but do not on their own provide activation (Abid Ali et al, 2025).
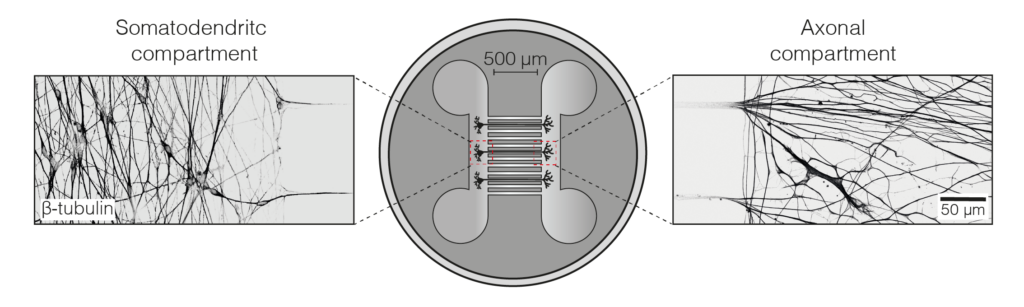
We set up an iNeuron system in our lab, allowing us to endogenously tag dynein and dynactin for near-single molecule imaging during axonal transport. We showed dynein can move the whole length of the axon (some 500 um) in one go. We could also follow their delivery, by kinesin motors, from the cell body out to the axon tip. To our surprise we found that a substantial fraction of dynein was transported out at a different speed from dynactin. This suggests that in the axon a bulk of dynein is transported outward separately from dynactin (Fellows et al, 2024).
Dynein cargos
We are increasingly focusing on how specific cargos are recognised and transported by the microtubule machinery.
The intracellular bacterium Orientia tsutsugamushi which causes scrub typhus is known to hijack dynein. In collaboration with Jeanne Salje (Cambridge), we found that bacterial autotransporter ScaC specifically binds dynein adaptors BICD1 and BICD2. A double knockout of both adaptors abolished clustering of O. tsutsugamushi at the nucleus demonstrating these adaptors are both necessary and sufficient for that stage of the bacterial lifecycle (Mannigrasso et al, 2025).
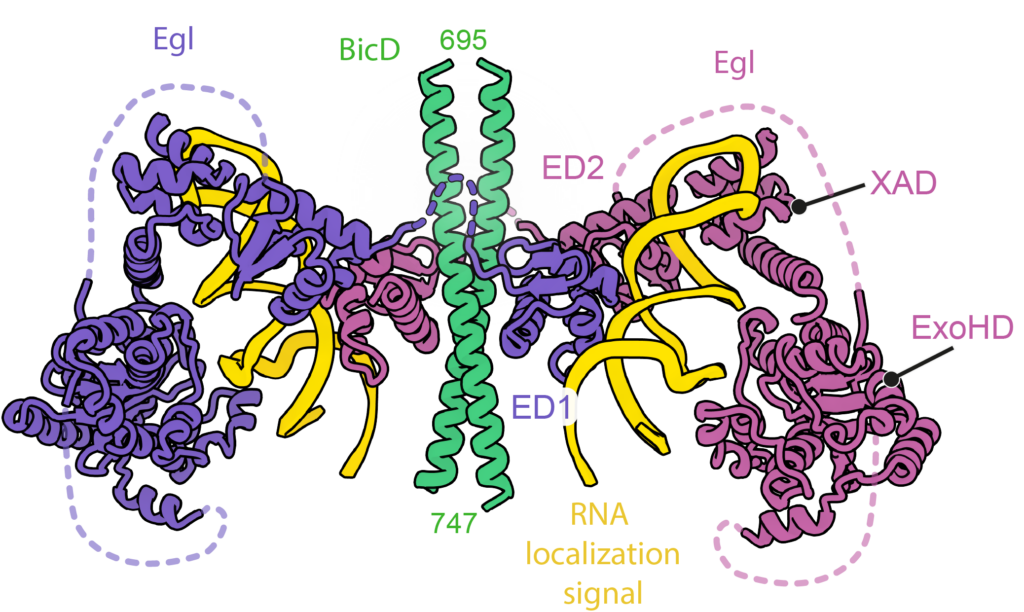
A classic model system for understanding mRNA localization is the Drosophila protein Egalitarian which binds to the dynein adaptor BicD. Localization signals in multiple mRNAs have been mapped but apart from being stem-loops have no obvious similarity in primary sequence or secondary structure. Working with Simon Bullock’s group (MRC LMB) we solved cryo-EM structures of 6 different mRNA localization signals bound to BicD-Egl. These identified the underlying patterns in the mRNAs and explained why they had been difficult to spot.
Axonemal Dyneins
Outer arm axonemal dyneins (ODAs) are the main drivers of cilia movement and are frequently mutated in human motile ciliopathies.
We identifying a novel factor required to deliver newly synthesised ODAs to Tetrahymena cilia (Maliet al., 2021). The factor, which we named Shulin, is conserved in humans, where, based on our work, it was named as an axonemal dynein assembly factor (DNAAF9).
Cryo-EM showed how Shulin binds and inhibits ODAs and we found that it accumulates in regenerating cilia, with its entry dependent on ODAs. This led to a model in which Shulin acts as an inhibitor that escorts ODAs into cilia and is released once they reach their destination. The work included the first structure of an axonemal dynein.
Follow-up studies are being carried out by Girish Mali in his own group at the University of Oxford.
Past Research
For past research highlights please click here