In vitro protein evolution and selection
Antibodies form a central pillar of the immune system. These remarkable molecules allow the immune system to recognise an effectively infinite array of antigens. The ability of antibody producing B cells to produce high affinity antibodies to a novel antigen within days of exposure is the result of the a series of mechanisms that specifically diversify the genes encoding the variable region of the antibody molecule.
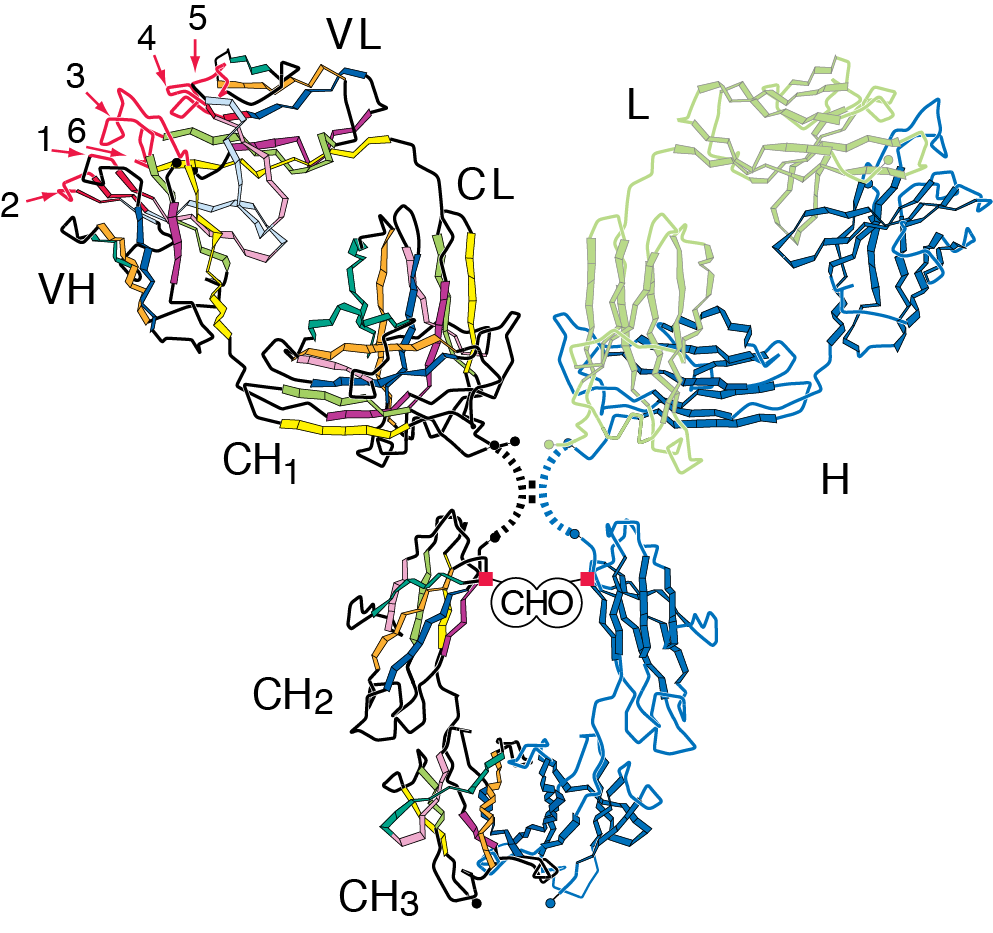
1. The antibody molecule
The antibody molecule (Figure 1) is made of two heavy and two light chains, indicated in blue and green respectively on the right hand side of the image. The two pairs of heavy and light chains are linked to each other by a disulphide bridge. The chains form a series of domains, a variable domain (VH) and three or four constant domains (CH1 - 3) depending on the isotype. The variable domain is responsible for recognising antigen. Contact with antigen is generally made through the hypervariable loops (three in VH and three in VL, indicated in red), which are encoded by the complementarity determining regions (CDRs) of the antibody genes. The conformation of these loops is also significantly influence by the sequence of the rest of the VH domain, which makes up the framework region. The constant regions specify the effector functions of the molecule by interfacing with components of the immune system such as macrophages, complement and natural killer cells.
Antibody diversity is produced in two stages (Figure 2). The first is independent of antigen and creates a 'naive' repertoire of specificities through a process called V(D)J recombination in which the exon encoding the variable region is created by recombination of gene segments, driven by the RAG proteins. The large repertoire that is created is able to bind a broad range of antigens, but usually only with low affinity. Following a productive encounter between and B cell and an antigen, the initially low affinity of the antibody is improved over the course of ten days to two weeks. This process of 'affinity maturation' results from a Darwinian mutation and selection process in which the variable segment of the antibody gene is diversified and variants with higher affinity selected to continue to divide and mutate.
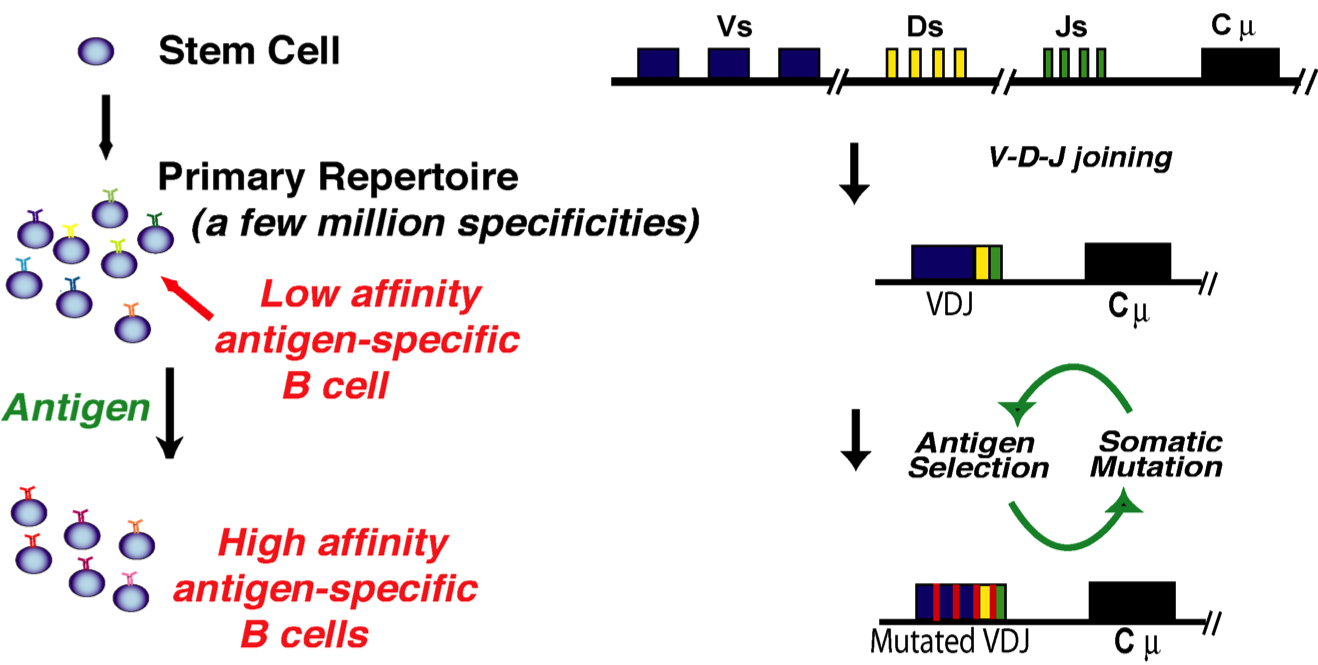
2. Primary and secondary antibody diversification
.
Antibody diversification is driven by the enzyme activation induced deaminase (AID). AID deaminates dC to dU in the immunoglobulin variable genes. The processing of this dU then leads to a series of mutagenic outcomes that diversify the sequence of the V gene (Figure 3). In many species, somatic hypermutation is accompanied, or indeed largely replaced, by gene conversion in which templated sequence changes are introduced into the variable region from an array of nearly pseudogenes.
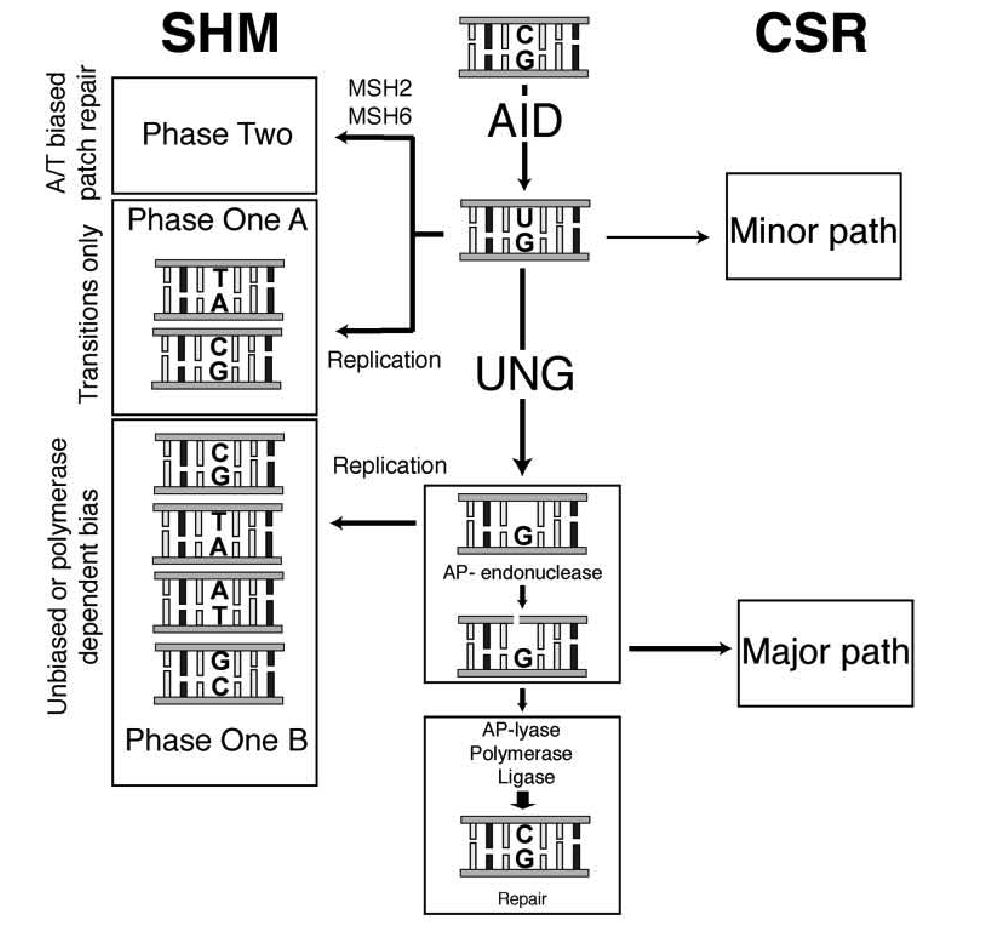
3. The Neuberger model of Ig diversification
A model of Ig diversification developed by the late Michael Neuberger illustrating the consequences of dC deamination in the Ig variable region in humans. The formation of dU creates a mismatch, which can be recognised by the mismatch repair proteins MSH2/MSH6. This leads to formation of gaps that are filled in by error-prone DNA synthesis that tends to generate mutations at A and T bases. Replication of dU, or the abasic site that results from its excision, generates mutations at G and C bases. (SHM = somatic hypermutation; CSR = class switch recombination, a process in which the isotype of the antibody molecule is changed.)
Transformed B cell lines that perform Ig diversification spontaneously during in vitro culture provide a very powerful tool for selection and evolution of novel antibodies against desired antigens. The chicken DT40 cell line has proved particularly useful because of its genetic tractability allowing its Ig variable genes to be manipulated or replaced. We have exploited this property to create starting libraries of surface-displayed single chain Fvs (truncated antibodies comprising just a single VH fused to a VL) that can be evolved and selected (Figure 4). The system can also be used to select and evolve non-antibody protein interactions. For example, while on sabbatical in the lab, Prof Nick Brindle (University of Leicester) created a novel ligand trap against Angiopoietin 2.
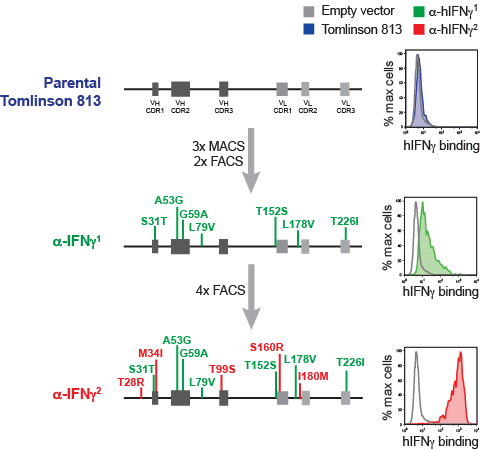
4. Example of the selection and evolution of a human scFv in DT40
A pool of DT40 cells expressing a mini-library of 11 different scFv was panned for cells binding to human IFN γ using a combination of magnetic bead sorting (MACS) and flow cytometry sorting (FACS). An initial binder (green) was subsequently matured (red) with 12 distinct point mutations separating this scFv from its parental sequence. For futher details see Lim et al. 2016.
We are continuing to develop and improve this system to create a facile platform that can be used to rapidly create or mature human antibody specificities against the widest possible range of antigens.