ost-translational modifications dynamically alter the molecular function, localization and stability of proteins, and regulate almost all aspects of biology. There is extensive crosstalk between modifications, making it challenging to address the function of any particular modification by genetic approaches. Spectacular progress has been made in defining the modified proteome, but on its own this provides little information about how post-translational modifications alter protein structure, function, activity or biological outcome.
Biochemistry, enzymology and structural biology provide a powerful set of approaches to address the role of a post-translational modifications, but it is often difficult to produce a homogeneously and site-specifically modified protein on which to perform studies (for many modifications that are identified in large screens the enzymes that carry out the modification in vivo are unknown, and even when they are known they often cannot be used to modify the protein site-specifically or completely in vitro).
We have reported genetic code expansion based methods for introducing post-translational modifications – including Tyr nitration, Lys acetylation, Lys monomethylation and dimethylation and Lys ubiquitylation – into recombinant proteins. Moreover, we have used the approaches we have developed to reveal the effect of the modifications on protein structure, function, activity and regulation, and thereby provided new insight into diverse areas of biology.
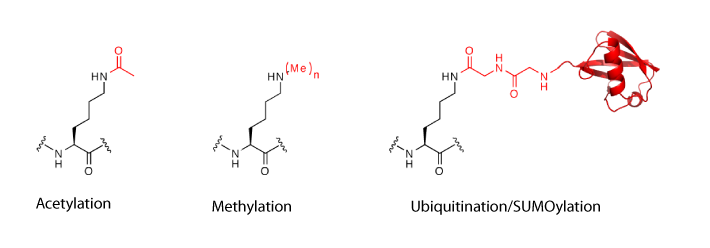
Installing Lys acetylation. Lys residues in proteins may be acetylated by acetyltransferases and deacetylated by histone deacetylases (HDACs) or sirtuins. Lys acetylation is a key post-translational modification in regulating chromatin structure and function and epigenetic phenomena. It is also now well established that acetylation is a key regulator of diverse biological processes, from metabolism to signalling and immunity.
We evolved a variant PylRS–tRNACUA pair that site specifically incorporates Acetyl-lysine into recombinant proteins produced in E. coli, in response to an amber codon. Using this approach, numerous recombinant proteins bearing homogeneous acetylation at a defined sites have been produced and we, and others have used this approach to provide new insights into the role of acetylation that would be challenging or impossible to achieve by other methods. Recently we have provided insights into the role of acetylation in regulating chromatin structure and function, metabolism and enzyme activity, immunosuppression, viral infection and tumour suppressor and transcription factor activity, and two examples of this work, much of which is collaborative, are described below.
Investigating the role of H3K56 acetylation in ‘designer chromatin’. Acetylation of histone H3 at Lys56 (H3K56) has been implicated in diverse biological processes, including the definition of epigenetic status and the regulation of transcription, DNA replication and DNA repair. To address the effects of H3K56 acetylation on chromatin structure and function we produced recombinant H3 bearing quantitative acetylation at Lys56 and then reconstituted histone octamers and assembled them into nucleosome core particles and nucleosomal arrays. This allowed us to test how H3K56 acetylation may control diverse phenomena by altering higher order chromatin structure; remodelling or repositioning nucleosomes; allowing nucleosome core particle ‘breathing’ or opening on DNA; or recruiting factors to this modification. Contrary to the prevailing ‘dogma’, we found that H3K56 acetylation did not measurably affect the chromatin compaction of nucleosomal arrays. Remodelling assays of single nucleosome movements on DNA revealed that acetylation of H3K56 caused only modest increases in repositioning of nucleosomes by SWI/SNF and RSC remodelling enzymes.
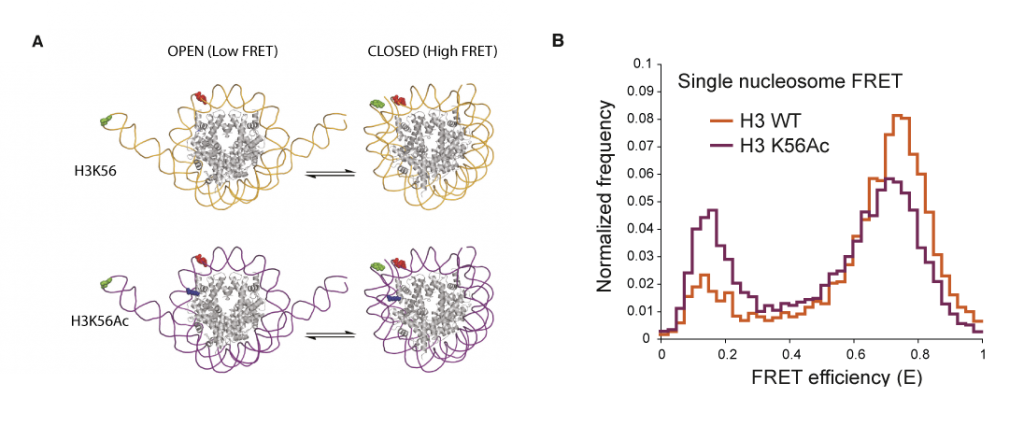
The crystal structure of the nucleosome core particle shows that H3K56 is near the entry and exit site of DNA and that Lys56 in H3 makes a water-mediated contact with the DNA phosphate backbone. Therefore, it was proposed that neutralization of the Lys charge by acetylation might alter DNA breathing through the local disengagement of regions of the DNA from the protein surface in the protein–DNA complex. This was experimentally demonstrated and quantified by single-nucleosome fluorescence resonance energy transfer (FRET) experiments. FRET probes on the DNA revealed an increase in breathing of DNA on the nucleosome core particles that contained acetylated H3K56 compared with those that contained wild-type H3. Moreover, by changing the position of the FRET probes on the DNA, it was possible to show that the acetylation-dependent increase in breathing localized to one turn of the DNA helix. This histone acetylation-dependent increase in DNA breathing may increase the accessibility of the DNA and the modified nucleosome to other factors that alter chromatin, providing a possible biophysical basis for observed cellular phenomena. The ability to produce homogeneously and quantitatively modified protein at Lys56 of histone H3 was key to making interpretable measurements in these single-molecule experiments.
CYPA acetylation during viral infection and immunity. Cyclophilin A (CYPA; also known as PPIA) is a cis–trans prolyl isomerase, and two of the most important roles of CYPA are in controlling immunosuppression and viral infection. CYPA is the target of the widely used immunosuppressive molecule cyclosporine: the complex formed by CYPA and cyclosporine binds and allosterically inhibits calcineurin, leading to suppression of the T cell-mediated immune response. CYPA is also an essential host protein for the efficient replication of several viruses, including HIV-1. During HIV-1 infection, CYPA interacts with the capsid protein GAG and is packaged into budding HIV-1 virions; and inhibition of CYPA by cyclosporine substantially decreases viral titre in human cells. It has been proposed that disassembly of the HIV capsid, which is an essential step in HIV infection, may be catalysed by CYPA-mediated isomerization of the Gly89–Pro90 bond in the HIV capsid.
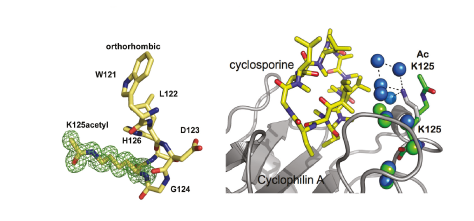
The acetylation of a cyclophilin at Lys125 was identified in a proteomics screen. We subsequently demonstrated that a substantial proportion of CYPA is acetylated in HeLa cells and human T cell lines, suggesting that the acetylated form of the protein is biologically relevant. To test this, recombinant CYPA bearing homogenous acetylation at Lys125 was prepared by overexpression in E. coli using an acetyllysyl-tRNA synthetase–tRNACUA pair, allowing detailed biophysical and enzymatic characterization of acetylated CYPA and associated complexes, as well as the first high-resolution structures of an acetylated protein and associated complexes. The acetylation of CYPA at Lys125 decreased its affinity for cyclosporine 20-fold. Comparison of the crystal structure of acetylated CYPA in complex with cyclosporine and the same complex for non-acetylated CYPA revealed that the acetylation of Lys125 disrupts a network of five water molecules that form an interface between unacetylated CYPA and cyclosporine. This acetylation-mediated solvent remodelling illustrates how acetylation may control protein–small molecule interactions without eliciting rearrangement of the structure or inducing allosteric changes. CYPA acetylation also decreased the affinity of the CYPA–cyclosporine complex for calcineurin twofold. As cyclosporine-mediated immunosuppression requires its binding to CYPA and the CYPA–cyclosporine complex binding to calcineurin, both of which are decreased by acetylation, the effect of CYPA acetylation on immunosuppression may be greater than the effect measured for each step individually.
Measurements on the CYPA–HIV-1 capsid interaction revealed that neither acetylation nor a Lys-to-Gln mutation at position 125 changes the affinity of CYPA for the HIV-1 capsid, but that the enthalpy and entropy contributions to this affinity do change. Therefore, the data are consistent with acetylated CYPA, Lys125Gln CYPA and wild-type CYPA binding the capsid through different modes, demonstrating that, although Gln is often used by researchers as a mimic of acetylation, it is not a good mimic for the acetylation of CYPA. The crystal structure of acetylated CYPA in complex with the HIV-1 M group capsid N-terminal domain also showed that the Gly89–Pro90 peptide bond of the HIV-1 capsid was in the cis form, rather than the trans form observed in the unacetylated CYPA complex, and acetylation decreased the catalytic efficiency for cis–trans prolyl isomerization of a model substrate 35-fold. These data are consistent with a model in which acetylation of CYPA decreases capsid isomerization and, therefore, potentially has an impact on capsid disassembly and HIV-1 infection. Together, these studies also demonstrate how insight into the consequences of a modification on biological function can be obtained through the synthesis and study of the modified protein.
Installing Lys ubiquitylation. The site-specific post-translational modification of target proteins with the 76-amino-acid protein ubiquitin regulates many aspects of eukaryotic cell biology. Ubiquitylation, in which the carboxy-terminal carboxylate of ubiquitin is attached to a Lys residue in a target protein to form an isopeptide bond, is naturally mediated by E1, E2 and E3 enzymes and removed by deubiquitinases (isopeptidases). However, the specific combinations of enzymes that are used to ubiquitylate many proteins are unknown, making it challenging to understand the roles of protein ubiquitylation and to decipher deubiquitinase specificity. As ubiquitin is itself a protein, this post-translational modification cannot be genetically encoded directly using an evolved synthetase and a tRNA, but we have developed methods that use genetic code expansion to site-specifically install Lys derivatives that allow ubiquitin to be attached to proteins through chemical steps that create a native isopeptide bond.
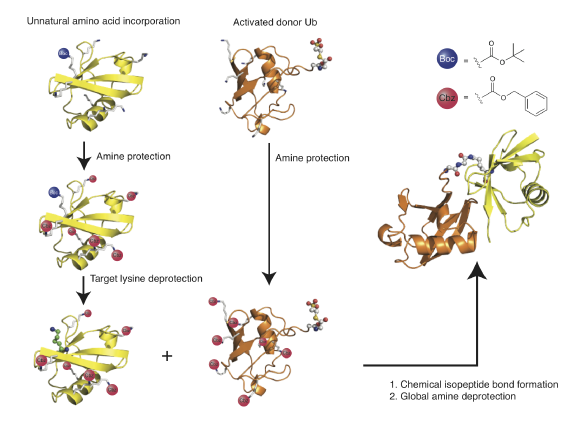
We have described two methods to create native isopeptide bonds between recombinant proteins and ubiquitin. In the first approach, termed GOPAL (genetically encoded orthogonal protection and activated ligation), we installed a protected version of Lys in response to the amber stop codon and defines the ultimate site of isopeptide bond formation. We then protected all other Lysines in the protein using an orthogonal protecting group and deprotected the amino acid installed in response to the amber stop codon, revealing a single free lysine amino group in the protein. We created a ubiquitin thioester by intein fusion thiolysis and protected all the lysines in ubiquitin. We formed the isopeptide bond between the ubiquitin thioester and the free amino group in the target protein via Silver catalysis and globally deprotected the product to reveal the ubiquitinated protein .
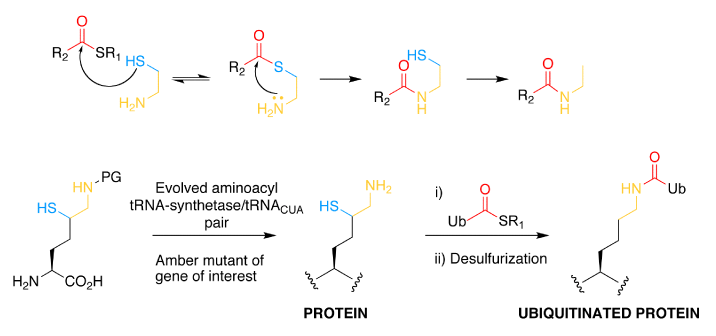
In a second approach, a PylRS–tRNA pair has been evolved to incorporate a δ-thiol-Lys derivative, which reacts with C-terminal thioesters of ubiquitin to yield an isopeptide bond. Desulphurization of the δ-thiol following isopeptide bond formation renders the approach entirely traceless. This approach was used to make ubiquitylated small ubiquitin-like modifier (SUMO), and simple extensions of the approach should allow access to large proteins bearing defined ubiquitylation, as well as to proteins bearing sumoylation or neddylation.
Atypical ubiquitin chain structure and deubiquitinase specificity. Atypical ubiquitin chains make up a large fraction of polyubiquitin in cells. However, the biological function, assembly and disassembly of atypical chains are less well understood than they are for Lys48 and Lys63 chains, which have established roles in proteasomal degradation and signalling, respectively. It was not previously possible to make these linkages so that their function and regulation could be investigated, as the specific ligases responsible for their synthesis were unknown. Using GOPAL, several atypical ubiquitin linkages were synthesized for the first time and this allowed the first structure of K6 linked ubiquitin to be determined and new deubiquitinase specificity to be revealed.
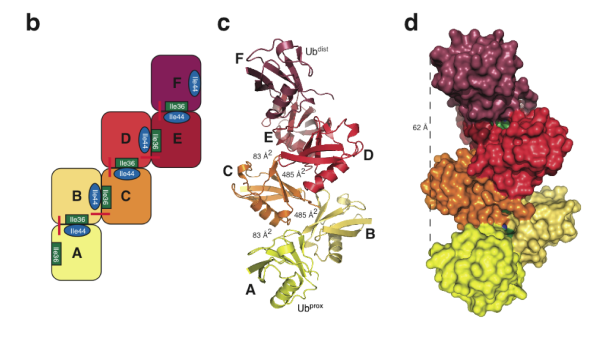
The crystal structure of Lys6-linked diubiquitin, synthesized by our methods, reveals an asymmetric compact conformation. The structure is distinct from previously observed diubiquitin structures for other isopeptide linkages and is also distinct from a computational model of Lys6-linked diubiquitin. Interestingly, iterative modelling of a longer Lys6-linked ubiquitin chain, on the basis of the diubiquitin structure, suggests the possibility that Lys6-linked ubiquitin may form a helical filament. Thus, the structure of Lys6-linked diubiquitin reveals molecular features that may subsequently explain how Lys6-linked ubiquitin polymers may be uniquely recognized in the cell to elicit particular biological outcomes.
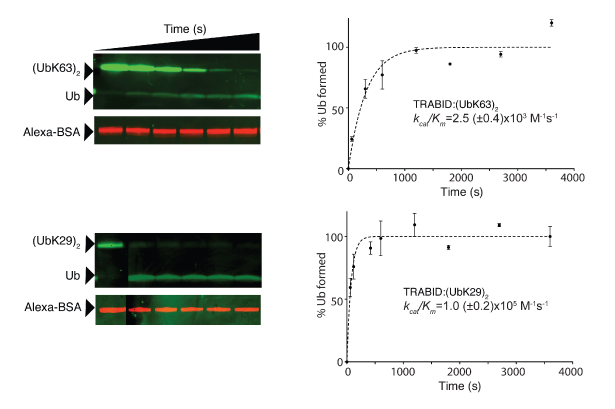
The ability to synthesize atypical ubiquitin chains allowed us to profile 10% of human deubiquitinases, which may each disassemble chains, for their isopeptide linkage specificity. These experiments revealed that atypical linkages are the preferred substrates of certain deubiquitinases; notably, the OTU domain deubiquitinase TRABID (also known as ZRANB1) cleaves Lys29-linked diubiquitin 40 times more rapidly than Lys63-linked ubiquitin95. As TRABID has been implicated as a positive regulator of WNT signalling, these experiments reveal a potential new link between WNT signalling and atypical ubiquitin chains and highlight the power of the approaches we have developed for assessing the regulation and biological roles of ubiquitinated proteins.
Installing Lys methylation. Lys residues in proteins can be monomethylated, dimethylated and trimethylated by methyltransferases, and methylation can be removed by demethylases. Both the degree and site of methylation may control biological outcomes, and the role of methylation on histones in regulating chromatin function has been extensively studied. For example, methylation of Lys9 in histone H3 (H3K9) is associated with heterochromatin formation.We demonstrated that monomethylated and dimethylated Lys can be installed into recombinant proteins.
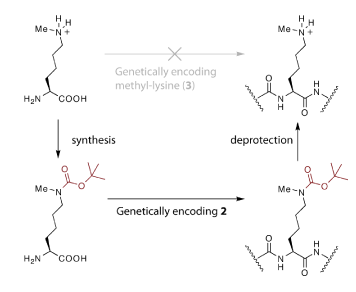
Using the PylRS–tRNACUA pair, a ‘protected’ version of monomethylated Lys (Nɛ-tert-butyloxycarbonyl-Nɛ-methyl-L-Lys) was site-specifically introduced into recombinant histone H3 produced in E. coli. Removal of the tert-butyloxycarbonyl group from the protein under mild conditions then revealed monomethylated Lys at the genetically programmed site. This two-step approach was necessary because Nɛ-methyl-L-Lys is so similar to Lys that it is difficult to create an aminoacyl-tRNA synthetase to specifically incorporate methylated Lys into proteins in the presence of the high Lys concentration that is constitutively present in the cell.
We have also described a technique for genetically directing Lys dimethylation in recombinant histones, thereby generating site-specifically dimethylated histones in large quantities. The approaches for installing monomethyl and dimethyl Lys in recombinant H3 have been used s to immunoprecipitate a known interaction partner (heterochromatin protein 1 (HP1)) of monomethylated and dimethylated H3K9.
References
DAXX envelops an H3.3–H4 dimer for H3.3-specific recognition.
Nature. 2012 491:560–565
S.J. Elsässer, H. Huang, P.W. Lewis, J.W. Chin, C.D. Allis, D.J. Patel
An Ankyrin-repeat ubiquitin-binding domain determines TRABID’s specificity for atypical ubiquitin chains.
Nature Struct. Mol. Biol. 2011 19:62-71
J.D.F. Licchesi, J. Miezczanek, T.E.T. Mevissen, T.J. Rutherford, M. Akutsu, S. Virdee, F.E. Oualid, J.W. Chin, H. Ovaa, M. Bienz, D. Komander.
Traceless & Site-Specific Ubiquitination of Recombinant Proteins.
J. Am. Chem. Soc. 2011 133:10708-10711
S. Virdee, P.B. Kapadnis, T. Elliott, K. Lang, J. Madrzak, D.P. Nguyen, L. Riechmann, & J.W. Chin.
A Dual Role of H4K16 Acetylation in the Establishment of Yeast Silent Chromatin.
EMBO J. 2011 30:2610-2621
M. Oppikofer, S. Kueng, F. Martino, S. Soeroes, S.M. Hancock, J.W. Chin, W. Fischle, S.M. Gasser.
Acetylation of Lysine 120 of p53 endows DNA-binding Specificity at Effective Physiological Salt Concentration.
Proc. Natl. Acad. Sci. U.S.A. 2011 108:8251-8256
E. Arbely, E. Natan, T. Brandt, M.D. Allen, D.B. Veprintsev, C.V. Robinson, J.W. Chin, A.C. Joerger, A.R. Fersht.
Molecular Basis for Ubiquitin and ISG15 Cross-reactivity in Viral Ovarian Tumor Domains.
Proc. Natl. Acad. Sci (USA) 2011 108: 2228-2233
M. Akutsu, Y. Ye, S. Virdee, J.W. Chin, D. Komander.
Engineered Diubiquitin Synthesis Reveals K29 Isopeptide Specificity of an OTU Deubiquitinase.
Nature Chem. Biol. 2010 6:750-757
S. Virdee, Y. Ye, D.P. Nguyen, D. Komander & J. W. Chin.
[Featured in Chem. Eng. News. Oct 11 2010, Chem. Eng. News (Highlights of the Year) Dec 20 2011., Angew. Chem. 2011 49:9042-9044]
Genetically Directing Ne, Ne dimethyl-L-lysine in Recombinant Histones.
Chem. Biol. 2010 17:1072-1076
D.P. Nguyen, M. Garcia-Alai, S. Virdee & J.W. Chin.
Regulation of Cellular Metabolism by Protein Lysine Acetylation.
Science. 2010. 237:1000–1004
22 co-authors
[Perspective in Science 2010 327: 964-965]
Acetylation Regulates Cyclophilin A Catalysis, Immunosuppression & HIV Isomerization.
Nature Chem. Biol. 2010 6:331-337
M. Lammers, H. Neumann, J.W. Chin, L. C. James.
[Featured in Chem. Eng. News. 12 April 2010 88:15 p28]
Genetically Encoding Ne-methyl-L-lysine in Recombinant Histones.
J. Am. Chem. Soc. 2009. 131:14194-14195
D. P. Nguyen, M. Garcia Alai, P.B. Kapadnis, H. Neumann, & J.W. Chin.
A Method for Genetically Installing Site-specific Acetylation in Recombinant Histones Defines the Effects of H3 K56 Acetylation.
Mol. Cell. 2009. 36:153-163
H. Neumann, S. Hancock, R. Buning, A. Routh, L. Chapman, J. Sommers, T. Owen-Hughes, J. van Noort, D. Rhodes, & J.W. Chin.
[Featured in Nature Chem. Biol. 2009. 5: 787; Chem. Biol. 2009. 16:1027-1029. Nature Methods 2009 6: 865]
Genetically Encoding Ne-acetyllysine in Recombinant Proteins.
Nature Chem. Biol. 2008. 4: 232-234.
H. Neumann, S.Y. Peak-Chew & J.W. Chin.
[Featured in Nature 2008. 45: 868, Chemistry & Engineering News (ACS). 2008. Feb 25, p38, Chemistry World (RSC), 2008. Feb 18]
Genetically Encoding Protein Oxidative Damage.
J. Am. Chem. Soc. 2008. 130: 4028-33
H. Neumann, J.L. Hazen, J. Weinstein, R. A. Mehl & J. W. Chin.
