dvanced imaging approaches can provide detailed information about the spatial and temporal organization of single proteins, as well as information about conformational dynamics and interactions of individual proteins. However, as imaging approaches advance, the information that can be gained about protein function in cells is increasingly limited by the precision with which (optical) probes can be attached to proteins in their native environment.
Current protein-labelling methods involve the use of fluorescent protein fusions self-labelling proteins (for example, SNAPtag, HALOtag and CLIPtag), ligases (such as biotin ligase, lipolic acid ligase, sortase and phosphopantetheinyl transferase) and self-labelling tags (for example, tetracysteine and tetraserine). Although some of these approaches allow rapid labelling and have had substantial impact on biological studies, they require the use of protein fusions and/or the introduction of additional sequences into the protein of interest. This may disturb the structure and function of the protein and can make it challenging to place probes at any position in a protein. Moreover, the range of probes that can be incorporated by some of these techniques is limited.
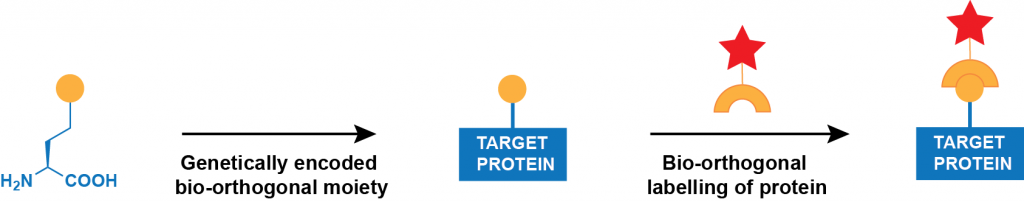
Genetic incorporation of an unnatural amino acid bearing a bio-orthogonal ‘handle’. Ideal methods for protein labelling would (i) allow probes to be placed easily at any position in any protein expressed in prokaryotic and eukaryotic cells and organisms, (ii) be rapid and quantitative, (iii) be specific for a user-defined site in a protein, (iv) require no toxic reagents and generate no toxic by-products, (v) show ‘turn-on’ fluorescence with minimal off-site or background labelling and (vi) allow for labelling with diverse probes. In principle, the genetically encoded, site-specific incorporation of unnatural amino acids that bear bioorthogonal functional groups would allow the labelling of specific proteins at defined sites with essentially any probe (Figure 1).
We demonstrated the genetic incorporation of amino acids bearing aliphatic azides and alkynes into recombinant proteins using the pyrrolysyl tRNA synthetase/tRNACUA pair, and the quantitative, site specific labeling of recombinant proteins bearing these amino acids with fluorophores and biotin in Copper (I) catalysed ‘click’ reactions. This approach has been used by our group, and adopted by others, for protein labeling and conjugation. However, the Copper (I) catalysed ‘click’ reactions are not generally applicable to labeling in live cells.
In the search for faster reactions for site specific protein labeling we evolved a pyrrolysyl tRNA synthetase/tRNACUA pair variant to genetically encode a set of 1,2 amino-thiol bearing amino acids. These amino acids react with cyanobenzothiazole-based probes. Using this approach we demonstrated, in combination with cysteine labeling, the labeling of a protein with two different probes. While this reaction allowed protein labeling in an hour with 10 equivalents of label, the 1,2 aminothiol formed thiazolidine adducts with aldehydes and ketones in cells, making it unsuitable for in vivo labeling. We have also investigated new bioorthogonal reactions, including the reaction of aryl-sydnones and strained alkynes and genetically encoded components of these reactions for protein labelling.
Inverse electron-demand Diels-Alder reactions. Bioorthogonal groups, including azides, alkynes, ketones, anilines, alkenes, tetrazoles and 1,2-aminothiols, have been genetically encoded using amber suppressor aminoacyl tRNA synthetase/tRNACUA pairs. For established reactions that have been demonstrated on proteins, the rate constants for the corresponding model reactions are in the range 10−2 M−1 s−1 to 10−4 M−1 s−1 (although for emerging approaches higher rates have been reported, Figure 2).
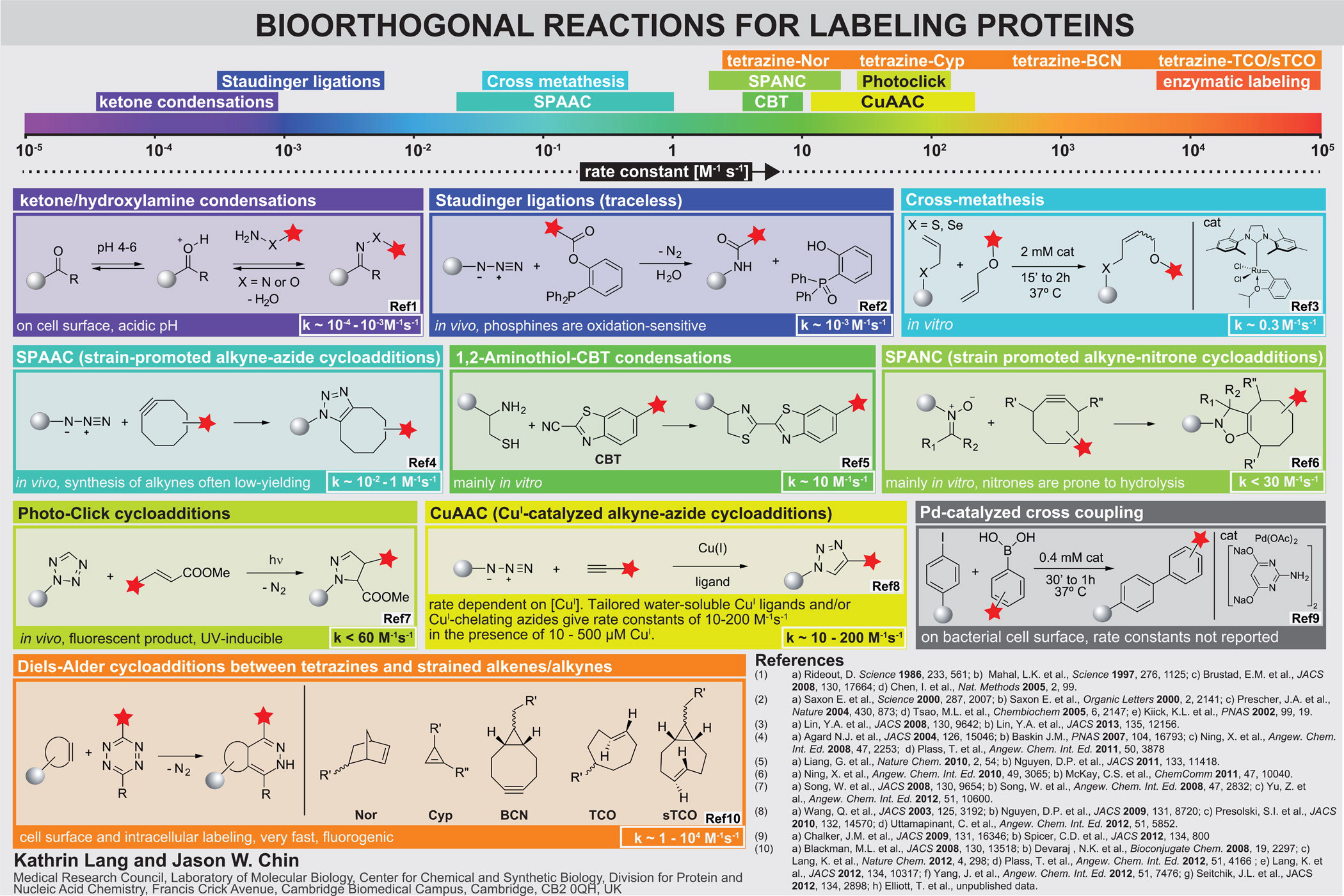
Several strained alkenes and alkynes (including norbornenes, trans-cylcooctenes and bicyclo[6.1.0]nonyne) react rapidly and specifically with tetrazines in inverse electron-demand Diels-Alder cycloadditions reactions to form stable adducts. These reactions occur at ambient temperatures in water with nitrogen gas as the only side product. The rate constants for these reactions are up to to 105 M−1 s−1, which is extraordinarily fast and approaches the rate of many enzyme mediated labelling procedures. Since the tetrazine quenches many red fluorophores to which it is attached, but the quenching is released upon reaction with strained alkenes or alkynes, these reactions are fluorogenic, minimizing background fluorescence in labelling reactions.
We have evolved aminoacyl-tRNA synthetase/tRNA pairs for the efficient site specific incorporation of several unnatural dienophile-bearing amino acids 1-3, bearing a norbornene, trans-cylcooctene and a bicyclo[6.1.0]nonyne Figure 3a) and well as a 1,3 di-substituted cyclopropene into proteins expressed in E. coli. We demonstrated the site-specific labelling of proteins in vitro and in E. coli with tetrazine-fluorophore conjugates in a Diels-Alder reaction with inverse electron demand (Figure 3b).
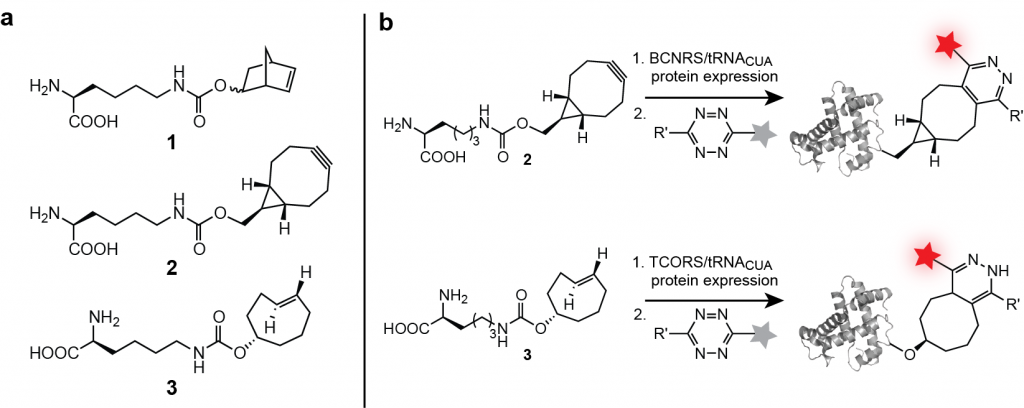
These reactions are very specific, rapid and the tetrazine fluorophores, which are initially weakly fluorescent become strongly fluorescent once attached to the protein via the chemical reaction, making the signal to noise of this labeling approach superior. We demonstrated that the labeling of these encoded unnatural amino acids 1–3 is specific with respect to the entire soluble E. coli proteome and several orders of magnitude faster than established bio-orthogonal reactions, allowing quantitative labeling of recombinant protein in seconds.
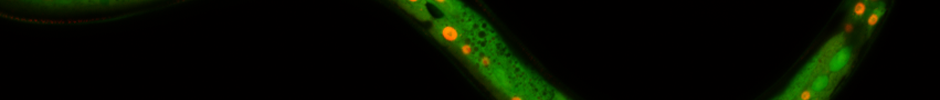
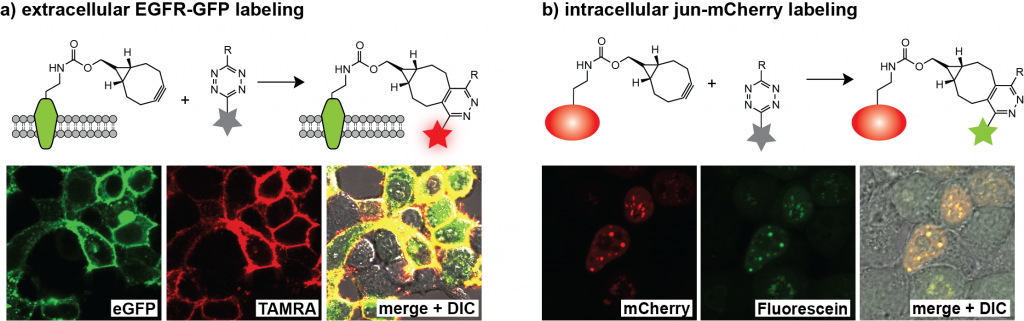
Site-specific protein labelling in live mammalian cells. We demonstrated that unnatural amino acids 1–3 can be encoded genetically with high efficiency into proteins in mammalian cells using the engineered aminoacyl-tRNA synthetase/tRNA pairs derivatives evolved for incorporation of amino acids 1–3. We demonstrated the first site specific protein labeling in live mammalian cells via genetic code expansion by expressing an epidermal growth factor receptor (EGFR)-GFP fusion bearing an amber codon in the extracellular portion of the receptor in HEK-293 cells containing the engineered aminoacyl-tRNA synthetase/tRNA pair cultured in the presence of one of the unnatural amino acids 1–3. When incubated with a tetrazine-tetramethylrhodamine (TAMRA) conjugate, clear labeling of cells cultured in the presence of amino acid 1–3, was observed within 2 minutes (the first time point we could measure) and TAMRA fluorescence clearly co-localized with cell-surface EGFR-GFP fluorescence (Figure 4a). In control experiments we observed neither labeling of cells bearing EGFR-GFP with another unnatural (non-tetrazine-reactive amino acid) amino acid nor nonspecific labeling of cells that did not express EGFR-GFP.
To demonstrate the rapid labeling of an intracellular protein in mammalian cells, we expressed a transcription factor, jun with a C-terminal mCherry fusion from a gene bearing an amber codon in the linker between JunB (jun) and mCherry. In the presence of amino acid 2 and the corresponding engineered aminoacyl-tRNA synthetase/tRNA pair (BCNRS/tRNACUA) the jun-2-mCherry protein was produced in HEK cells and, as expected, localized in the nuclei of cells. Labeling with a cell-permeable tetrazine-diacetylfluorescein conjugate resulted in green fluorescence that co-localized nicely with the mCherry signal at the first time point analyzed (Figure 4b). No specific labeling was observed in non-transfected cells in the same sample or in controls cells expressing jun-mCherry with a non-tetrazine-reactive amino acid, further confirming the specificity of intracellular labeling.
We have demonstrated the efficient site-specific incorporation of unnatural amino acids 1–3 into proteins in E. coli and mammalian cells and their efficient, specific, fluorogenic, and rapid labeling with tetrazine fluorophore conjugates. These experiments demonstrate that is possible to harness the intrinsic reactivity of small molecules to achieve labeling rates comparable to those facilitated by enzymatic catalysis. We have demonstrated the advantages of genetically encoding components of the inverse electron demand Diels Alder reaction for labelling proteins in vitro, in E. coli, in live mammalian cells, in C. elegans and Drosophila melanogaster.
References
Conformationally strained trans-cyclooctene with improved stability and excellent reactivity in tetrazine ligation.
Chem. Sci. 2014 DOI:10.1039/c4sc01348d
A. Darko, S. Wallace, O. Dmitrenko, M.M. Machovina, R. A. Mehl, J. W. Chin, J.M. Fox.
Concerted, Rapid, Quantitative, and Site-specifc Dual Labeling of Proteins
J. Am. Chem. Soc. 2014 DOI:10.1021/ja4129789
A. Sachdeva, K. Wang. T. Elliott, J.W. Chin.
Optimized orthogonal translation of unnatural amino acids enables spontaneous protein double labeling and FRET
Nature Chemistry. 2014 6:393-403.
K. Wang, A. Sachdeva, D. J. Cox, N. M. Wilf, K. Lang, S. Wallace, R. A. Mehl, J. W. Chin.
[Featured in News & Views Nature Chem. 2014 6:379-381]
Proteome Labeling and Protein Identification in Specific Tissues at Specific Developmental Stages in an Animal
Nature Biotech.2014 32:465-472.
T.S. Elliott, F.M. Townsley, A. Bianco, R.J. Ernst, A. Sachdeva, S.J. Elsaesser, L. Davis, K. Lang, R. Pisa, S. Greiss, K.S. Lilley, J.W. Chin.
[Featured in News & Views, Nature Biotech. 2014 32:445-446, Spotlight, ACS Chem. Biol. 2014 9:1068]
Cellular Incorporation of Unnatural Amino Acids and Bioorthogonal Labeling of Proteins
Chem. Rev. 2014 114:4764-4806.
K. Lang, J.W. Chin.
Strain promoted sydnone bicyclo-[6.1.0]-nonyne cycloaddition
Chem Sci. 2014 5:1742-1744.
S. Wallace, J.W. Chin.
Bioorthogonal reactions for labeling proteins
ACS Chem. Biol. 2014 9:16-20.
K. Lang, J.W. Chin.
Genetic Encoding of Bicyclononynes and trans-Cyclooctenes for Rapid Site-Specific Protein Labeling in Vitro and in Live Mammalian Cells via Fluorogenic Diels-Alder Reactions.
J. Am. Chem. Soc. 2012 134:10317-10320
K. Lang, L. Davis, S. Wallace, M. Mahesh, D. J. Cox, M. L. Blackman, J. M. Fox & J.W. Chin.
Genetically encoded norbornene directs site-specific cellular protein labelling via a rapid bioorthogonal cycloaddition.
Nature Chem. 2012 4:298-304.
K. Lang, L. Davis, A Deiters, J.W. Chin.
[Featured in Nature Chem. 2012 4: 248-250]
Genetically Encoded 1,2 Aminothiols Facilitate Rapid and Site Specific Protein Labeling via a Bio-orthogonal Cyanobenzothiazole Condensation.
J. Am. Chem. Soc. 2011 133:11418-11421
D.P. Nguyen, T. Elliott, M. Holt, T.W. Muir, & J.W. Chin.
[Featured in ACS Chem Biol. 2011 8: 768-770]
Genetic Encoding and Labeling of Aliphatic Azides and Alkynes in Recombinant Proteins via a Pyrrolysyl-tRNA Synthetase/tRNACUA Pair and Click Chemistry.
J. Am. Chem. Soc. 2009. 131: 8720-8721
D. P. Nguyen, H. Lusic, P.B. Kapadnis, A. Deiters & J.W. Chin.