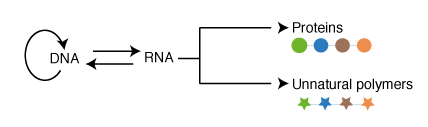
hile cellular translation on the ribosome normally makes proteins composed of the natural 20 amino acids, it represents the ultimate paradigm for the encoded, and evolvable, synthesis of proteins containing additional amino acids, and ultimately the synthesis and evolution of completely unnatural polymers. Since the ribosome uses tRNA adapter molecules it makes polymers that are chemically independent of the template; this should allow distinct polymers to be made without interfering with their encoding. Since the ribosome uses a single set of active sites for bond formation, and couples these to a translocation activity it can make very long polymers with very high fidelity.
There are several major challenges in genetically encoding multiple unnatural amino acids into proteins in living cells. First, unique new codons are required that can be used to encode the incorporation of unnatural amino acids at specific sites in proteins. Since the 64 triplet codons are used in the genome of most organisms for encoding natural proteins additional codons (such as quadruplet codons) might be used to encode the incorporation of unnatural amino acids. Second, new aminoacyl-tRNA synthetase tRNA pairs that are orthogonal to the aminoacyl-tRNA synthetase/tRNA pairs in the host organism, and that uniquely direct the incorporation of an unnatural amino acid in response to a unique codon, are required. Finally the scope of cellular protein translation is limited to alpha-L amino acids and their close analogs, and alteration of the ribosome and potentially other components of the translational machinery are required to increase the chemical scope of translation.
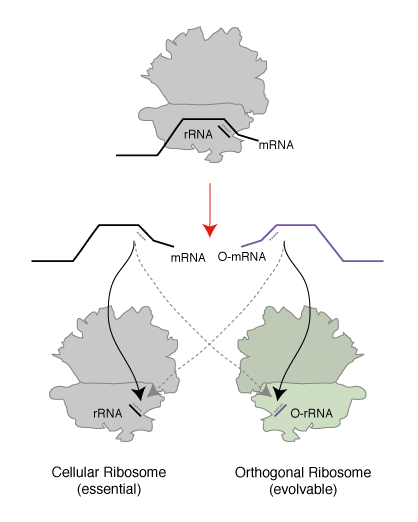
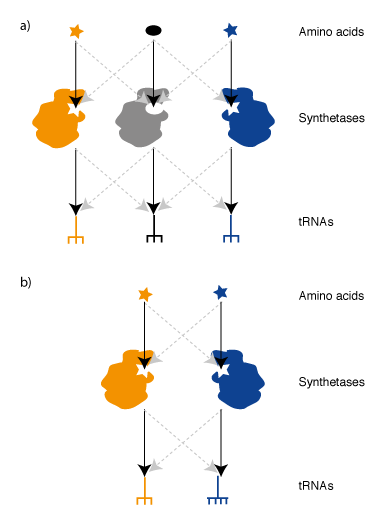
We realized that since the genetic code is a correspondence between amino acids and codons, which is set by the translational machinery, we could create a parallel genetic code (Figure 1) in the cell in two steps. In the first step we created an orthogonal ribosome that is selectively directed to a new orthogonal message that is not read by endogenous ribosomes. This creates a ribosome that, unlike the natural ribosome (which is required to synthesize the proteome and keep the cell alive and is therefore refractory to alteration of its sequence and structure), can be altered by synthetic evolution in the laboratory. The new orthogonal ribosome provides a platform for engineering the functional centres of the ribosome, that have been revealed by years of biochemistry and recent stunning advances in structural biology but which it has not been possible to alter to perform new functions. In a second step we altered the orthogonal ribosome to efficiently decode tRNAs on the orthogonal message that are not efficiently decoded on cellular messages. This is achieved by creating a bump on the tRNA (using an extended anticodon tRNA) and a corresponding hole in the orthogonal ribosome to accommodate the expanded tRNA. This approach allows the efficiency of the quadruplet decoding to be selectively enhanced on the orthogonal message, without affecting the decoding of cellular messages by natural ribosomes, and therefore enhances the efficiency of quadruplet decoding without creating toxic misreading of the proteome.
Creating Orthogonal Ribosome-mRNA Pairs. To create an orthogonal ribosome-mRNA pair in E. coli that operates in parallel with, but independent of, the natural ribosome we took advantage of the fact that the E. coli ribosome selects its cognate mRNA in the, rate determining, initiation stage of protein translation via a Shine Dalgarno sequence 7 to 12 bases 5’ of the AUG initiation codon. Genomic Shine Dalgarno sequences show substantial sequence variation, but the consensus sequence (AGGAGG) is complementary to the 3’ end of the 16S rRNA in the small subunit of the ribosome (CCUCCU). We created a large library of alternative Shine Dalgarno sequences that covers all combinations of nucleotides from 7 to 12 upstream to the AUG start codon in the mRNA. This allowed the alternative Shine Dalgarno sequences to vary in their sequence and spacing with respect to the start codon. We placed these sequences upstream of a gene encoding a fusion between cat (chloramphenicol acetyl-transferase) and upp (uracil phosphoribosyltransferase), which allows either positive or negative selection on the expression of the gene fusion upon the addition of distinct small molecules. In the presence of chloramphenicol the cat gene allows the selection of functional sequences on chloramphenicol and in the presence of 5-fluorouracil (5FU) the upp gene allows selection of non-functional sequences.
We first selected, on 5FU, for sequences upstream of the cat-upp fusion that do not allow the gene to be translated by the cell’s endogenous ribosome, creating a pool of potentially orthogonal ribosome binding sites. Next we created a library, based on the gene for the rrnB ribosomal RNA operon. This library allows cells to produce ribosomes that contain all possible combinations of nucleotides at the 3’ end of 16S rRNA. Growth of cells in the presence of chloramphenicol allows selection of orthogonal ribosomes that specifically translate the orthogonal messages. From the 109 combinations of potential ribosome mRNA pairs interrogated in this two step selection we found three classes of orthogonal ribosome-orthogonal mRNA pairs. These pairs differ in the base paired sequence by which the ribosome binds to its cognate message (Figure 2).
Evolving an Orthogonal Ribosome for Quadruplet Decoding. With orthogonal ribosome-mRNA pairs in hand we set out to evolve an orthogonal ribosome to efficiently decode quadruplet codons, which are poorly decoded on the natural ribosome. To create a ribosome that decodes quadruplet codons we first designed and created 13 structurally guided libraries in the decoding centre in the A-site of the ribosome, which is responsible for the fidelity of triplet decoding. Each library contains approximately 108 members, and together the libraries cover a molecular surface defined by 127 nucleotides of 16S rRNA. To select for orthogonal ribosomes that efficiently decode extended anticodon tRNAs we combined the library of orthogonal ribosomes with an extended anticodon tRNA that is aminoacylated by the E. coli seryl-tRNA synthetase, and incorporates serine in response to the AAGA quadruplet codon. Cells also contained an orthogonal message bearing a cat gene with an in-frame AAGA codon. Reading of the AAGA codon on mutant orthogonal ribosomes, using the extended anticodon tRNA, led to synthesis of the full-length chloramphenicol acetyl transferase protein. This allowed the selection, on chloramphenicol, of cells bearing orthogonal ribosomes that are able to facilitate the decoding of the extended anticodon tRNA (Figure 3).
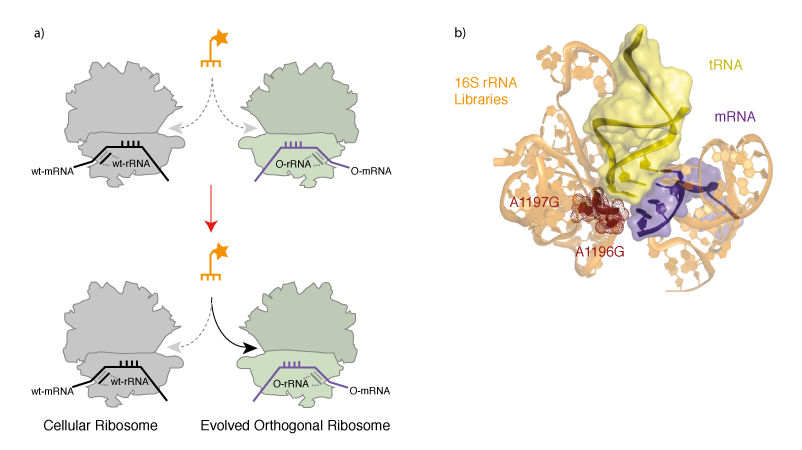
We selected a new orthogonal ribosome Ribo-Q1 that was able to efficiently decode a range of different quadruplet codons using extended anticodon tRNAs. Indeed, the levels of chloramphenicol resistance in cells bearing Ribo-Q1, a gene encoding cat with a AGGA codon, and the corresponding extended anticodon tRNA that is aminoacyated by E. coli seryl tRNA synthetase was comparable to the levels of resistance in cells bearing a wild type cat gene. This suggests that the level of quadruplet decoding on the evolved ribosome, in the presence of an efficiently aminoacylated tRNA, can approach the level of triplet decoding on the natural ribosome. Ribo-Q1 was derived from Ribo-X (an evolved orthogonal ribosome selected for enhanced amber suppression that allowed the first efficient and quantitative incorporation of multiple identical unnatural amino acids at specific sites) and was additionally able to efficiently decode the amber codon on an orthogonal message using amber suppressor tRNAs. Although we mutated 127 nucleotides in the decoding centre, the mutant ribosome contains two mutations with respect to Ribo-X. Mapping the selected mutations onto a crystal structure of the ribosome bound to a tRNA (Figure 4b) reveals that the selected mutations cluster where the 4th base of the extended anticodon might be expected to reside. This suggests that the selected mutations may result in a local rearrangement of ribosomal RNA structure. We demonstrated that Ribo-Q had excellent translational fidelity. Moreover we were able to efficiently decode several distinct quadruplet codons using tRNAs bearing distinct extended anticodons and Ribo-Q. This work therefore provides several blank codons that can be specifically decoded on the orthogonal message by Ribo-Q, and potentially assigned to new amino acids.
Synthetases & tRNAs for an Orthogonal Genetic Code. In order to take advantage of Ribo-Q for incorporating multiple distinct unnatural amino acids we require an orthogonal synthetase/tRNA pair for each new codon. Two useful aminoacyl tRNA synthetase pairs that are orthogonal in E. coli, the MjTyrS/tRNACUA pair and PylRS/tRNACUA pair from Methanosarcina species,have been discovered by import of the pairs from archaea (Figure 4). This approach that takes advantage of the fact that, while the genetic code is near universally conserved between organisms, the identity elements by which these archaeal synthetases recognize their cognate tRNA have fortuitously diverged, over the timescale of natural evolution, to create orthogonal pairs.The active site of MjTyrRS has been evolved in the laboratory to allow the incorporation of a range of aromatic unnatural amino acid variants. The PylRS/tRNACUA pair is a recent discovery in certain methanogens that naturally recognizes pyrrolysine and incorporates this amino acid in response to the amber stop codon. Unlike the pathway for incorporating selenocysteine, a synthetase/tRNA and amino acid are sufficient to direct the incorporation of pyrrolysine in response to the amber stop codon. Because the pyrrolysyl-tRNA synthetase/tRNACUA pair is orthogonal with respect to E. coli synthetases and tRNAs and does not use the natural 20 amino acids it has been possible to use this pair and its synthetically evolved variants to incorporate diverse unnatural amino acids into proteins.
We demonstrated that the non-cognate MjTyrRS /Methanosarcina barrkeri (Mb)tRNACUA and MbPylRS /Mj tRNACUA pairs do not function as amber suppressors, suggesting that the MjTyRS/tRNACUA pair and MbPylRS/tRNACUA pairs are mutually orthogonal in their aminoacylation specificities. To use these synthetases together to direct the incorporation of distinct amino acids in response to distinct codons we needed to differentiate the codons they decode. To alter the MjTyrRS/tRNACUA pair so that it incorporates an unnatural amino acid in response to a unique codon we first mutated the anticodon of the MjtRNACUA to UCCU, creating MjtRNAUCCU. Since MjTyrRS recognizes the anticodon this alteration produced a non-functional synthetase/tRNA pair. To create a synthetase that functions with MjtRNAUCCU to incorporate a useful unnatural amino acid in response to a quadruplet codon, we swapped MjTyrRS for MjAzPheRS ( a synthetase variant that recognizes p-azido-L-phenylalanine), and created a library of mutants in the region of the synthetase that recognize the anticodon. We then selected variant MjAzPheRS/tRNAUCCU pairs that can incorporate p-azido-L-phenylalanine in response to the AGGA codon (Figure 4b).
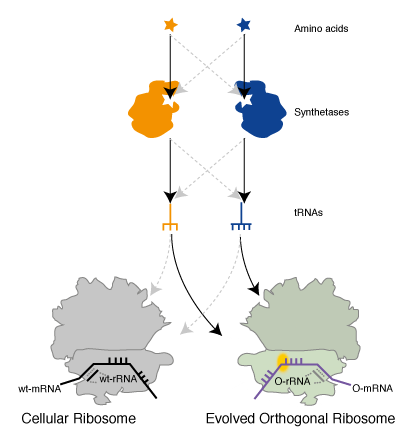
Genetically Encoding Emergent Properties in Proteins via Orthogonal Translation. To put together an orthogonal translation pathway for site specifically incorporating two unnatural amino acids we combined Ribo-Q with an mRNA containing a UAG and AGGA codon, MjAzPheRS/tRNAUCCU, and MbPylRS/tRNACUA. We demonstrated that full-length protein was generated only in the presence of p-azido-L- phenylalanine and an aliphatic alkyne, that we had previously shown to be a good substrate for PylRS, and that both amino acids were incorporated at the genetically encoded sites as judged by mass spectrometry. This experiment demonstrated the creation of a parallel pathway in the cells that incorporates unnatural amino acids in response to amber and quadruplet codons (Figure 5).
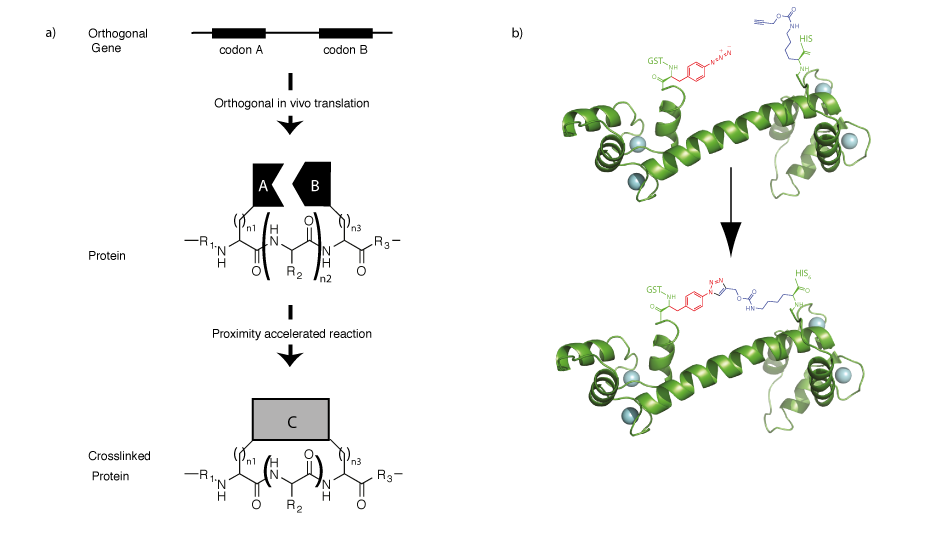
The ability to direct two unnatural amino acids into recombinant proteins allows us to program properties into proteins that are not a property of either amino acid individually but emerge from the interaction between the two amino acids (Figure 6). We demonstrated that by encoding an aliphatic alkyne and an azide containing amino acid it is possible to genetically program a rapid, proximity accelerated cycloaddition between these bioorthogonal groups to form a nanoscale, redox insensitive, triazole crosslink in a protein. Extensions of this approach may allow us to rapidly explore all possible crosslinks in proteins, and this approach may find utility in trapping particular functional states of proteins or in stabilizing protein therapeutics.
Since the synthetases derived from MjTyrRS and PylRShave each been used to encode numerous unnatural amino acids, it is now be possible to encode several hundred pairwise combinations of unnatural amino acids into proteins by simple extensions of our approach. By encoding new combinations of unnatural amino acids new properties, such as fluorescence, may be programmed into proteins, and this may facilitate the labelling of specific proteins in vivo
De novo Evolution of New Orthogonal Aminoacyl-tRNA Synthetase/tRNA Pairs. Ribo-Q provides numerous additional codons on the orthogonal mRNA. However, since only two orthogonal synthetase/tRNA pairs exist that can be used to incorporate distinct amino acids, only two distinct unnatural amino acids can be incorporated into a protein in the cell. A clear challenge in going from incorporating two unnatural amino acids to the synthesis of completely unnatural polymers is therefore to discover or invent strategies for generating new orthogonal aminoacyl-tRNA synthetase/tRNA pairs that can be used to encode additional unnatural amino acids into a single polypeptide.
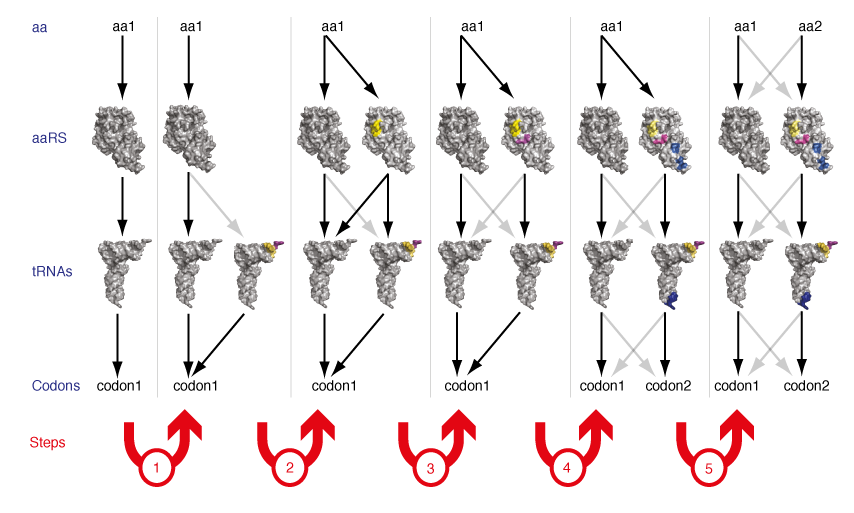
The two existing orthogonal synthetase tRNA pairs in E. coli were derived by import from heterologous organisms, taking advantage of the fact that while the genetic code is near universally conserved between known organisms the sequences and structures of synthetases and tRNAs have diverged through evolution. However, it is unclear how many mutually orthogonal synthetase/tRNA combinations can be discovered by taking advantage of natural evolutionary divergence. Moreover since the evolutionary record suggests that the current set of synthetases and tRNAs arose by gene duplication and specialization from a simpler basis set (for example tyrosyl-tRNA synthetase and tryptophanyl-tRNA synthetase appear to be derived from a common ancestor), we realized that it might be possible to extend this evolutionary process in the laboratory to generate new orthogonal synthetases and tRNAs de novo. We demonstrated that by a series of genetic selections on structurally targeted libraries in a synthetase/tRNA pair it is possible to evolve a new synthetase/tRNA pair that is orthogonal to both the synthetase/tRNA pair from which it was evolved and every other synthetase and tRNA in the cell (Figure 7). This work demonstrates that the small number of orthogonal synthetase/tRNA pairs that have been discovered in nature do not intrinsically limit the potential of genetic code expansion for encoding additional unnatural amino acids.
References
Concerted, Rapid, Quantitative, and Site-specifc Dual Labeling of Proteins
J. Am. Chem. Soc. 2014 DOI:10.1021/ja4129789
A. Sachdeva, K. Wang. T. Elliott, J.W. Chin.
Optimized orthogonal translation of unnatural amino acids enables spontaneous protein double labeling and FRET
Nature Chemistry. 2014 6:393-403.
K. Wang, A. Sachdeva, D. J. Cox, N. M. Wilf, K. Lang, S. Wallace, R. A. Mehl, J. W. Chin.
[Featured in News & Views Nature Chem. 2014 6:379-381]
Encoding Multiple Unnatural Amino Acids via a Quadruplet Decoding Ribosome.
Nature. 2010 468:441-444.
H. Neumann, K. Wang, L. Davis, M. Garcia-Alai, & J.W. Chin.
[Featured in Nature Meths 2010 7: 343, New Scientist 17 Feb 2010, Chem. Eng. News 22 Feb 2010 v88: 8, p9, National Public Radio (USA) Morning Edition, 15 Feb 2010]
De Novo Generation of Mutually Orthogonal Aminoacyl-tRNA synthetase/tRNA pairs.
J. Am. Chem. Soc. 2010. 132:2142-2144
H. Neumann, A. Slusarczyk, J.W. Chin.
[Featured in New Scientist 17 Feb 2010. Chem. Eng. News 22 Feb 2010 88: 8, p9]
Evolved Orthogonal Ribosome Purification for in vitro Characterization.
Nucleic. Acids Res. 2010 38:2682-2691
O. P. T. Barrett & J. W. Chin.
Synthesis of Orthogonal Transcription-Translation Networks.
Proc. Natl. Acad. Sci. U.S.A. 2009. 106: 8477-8482.
W. An & J.W. Chin.
[Featured in Nature 2010 463:288-290]
Evolved Orthogonal Ribosomes Enhance the Efficiency of Synthetic Genetic Code Expansion.
Nature Biotech. 2007. 25: 770-777.
K. Wang, H. Neumann, J. W. Chin.
[News & Views Nature Biotech. 2007. 25: 745-746.]
Functional Epitopes at the Ribosome Subunit Interface.
Nature Chem. Biol. 2006. 2: 254-258.
O. Rackham, K. Wang & J. W. Chin.
[News & Views, Nature Chem. Biol. 2006. 2: 231-232]
Synthesizing Cellular Networks from Evolved Ribosome•mRNA Pairs.
Biochem. Soc. Trans. 2006. 34: 328-329.
O. Rackham & J. W. Chin.
Cellular Logic with Orthogonal Ribosomes.
J. Am. Chem. Soc. 2005. 127: 17584-17585.
O. Rackham & J. W. Chin.
[Featured in Science 2005. 310: 1587]
A Network of Orthogonal Ribosome•mRNA Pairs.
Nature Chem. Biol. 2005. 1: 159-166.
O. Rackham & J. W. Chin.
[Featured in Nature 2005. 436: 306-307; Science 2005. 310: 1587; Chemistry & Engineering News, 2005, July 25, p44; Nature Methods 2005. 2: 640]