The Wnt signalling pathway is an ancient cell communication pathway that has important roles in development and cancer. Wnt signals elicit context-dependent transcriptional responses by stabilising a cytoplasmic effector called beta-catenin. This controls the embryonic development of tissues and organs in all animals, from the most primitive ones all the way to humans. In addition, Wnt/beta-catenin signalling is also active in the stem cells of virtually all tissues in grown-up animals, to control their self-renewal upon injury or wear-and-tear. If inappropriately activated, beta-catenin can cause cancer in multiple human tissues. Indeed, more than 90% of all colorectal cancers are thought to be initiated by mutational activation of beta-catenin.
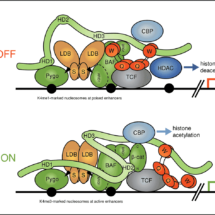
Beta-catenin binds to transcription (TCF) factors at Wnt-responsive DNA control regions of genes (called ‘enhancers’), to switch on gene expression programmes. These transcriptional switches rely on a multi-protein complex, the Wnt enhanceosome, which is tethered to DNA via TCF factors. Work By Mariann Bienz’s group, in the LMB’s PNAC Division, had previously proposed that the Wnt enhanceosome is made Wnt-responsive by a large protein called Legless (in the fruit fly Drosophila), or BCL9 and its close relative B9L (in other animals and humans). Legless/BCL9 proteins use one of their short conserved elements that are separated by long unstructured linker sequences, in ‘beads-on-a-string’ fashion, to dock to the chromatin-binding Pygo component of the Wnt enhancesome and another to capture the incoming beta-catenin, bringing it to TCF. Mariann’s group have now studied and determined the role of Legless/BCL9 within the enhancesome.
In order to probe the composition of the enhancesome, Moore (Laurens M.) van Tienen, from Mariann’s group, applied a relatively new technique called BioID proximity labelling, in order to define the proteins associated with BCL9/B9L in a human cell. BioID involves labelling all the proteins that a particular protein of interest comes into close contact with in live cells. Moore also applied CRIPSR/Cas9 genome engineering to Drosophila and human embryonic kidney (HEK) cells, to generate specific deletion mutants of Legless and BCL9/B9L that lack specific conserved elements, which allowed him to test the functional relevance of these elements during fly development and in human cells. The work was supported by the LMB’s Mass Spectrometry team, led by Mark Skehel, and the Flow Cytometry facility, led by Maria Daly, as well as technical advice and discussions with LMB group leaders, Simon Bullock and Mark van Breugel.
Mariann’s group discovered that BCL9/B9L binds to two distinct core components of the Wnt enhanceosome (in addition to Pygo) via distinct elements in their C-termini. By deleting these elements in Drosophila and HEK cells, they were able to demonstrate the functional relevance of these contact points to the Wnt enhanceosome in both systems. This corroborated and further refined the Wnt enhanceosome model, and identified BCL9/B9L as an integral component and scaffold of this complex, even when this complex is inactive in cells that are not stimulated by Wnt signals. Evidently, the Wnt enhanceosome is the key switch module that allows cells to translate an incoming Wnt signal into a transcriptional programme that determines the normal development and self-renewal of tissues, and that can also cause cancer if inadvertently activated. An unexpected discovery of this study was that BCL9/B9L and Pygo proteins are in close contact with nuclear co-receptor complexes, which has opened up the question whether these proteins are required to integrate Wnt/beta-catenin signalling with nuclear hormone receptor signalling. If so, this would have important implications for cancers that are driven by nuclear hormone receptors such as those of the breast and prostate. It could open up new avenues for therapeutic interventions.
The work was funded by the MRC and Cancer Research UK.