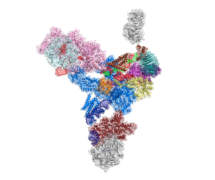
 Genes in eukaryotic organisms are frequently interrupted by non-coding segments called introns. Before the protein can be produced, the entire length of each gene including the introns, is transcribed to produce precursor messenger RNA (pre-mRNA). An immense and dynamic molecular machine known as the spliceosome then removes the introns and splices the coding segments together to form mature mRNA, which can then be translated into proteins. A joint team from Kiyoshi Nagai’s and Sjors Scheres’ groups in the LMB’s Structural Studies Division has used electron cryo-microscopy (cryo-EM) to reveal the detailed organisation of the spliceosome.
Genes in eukaryotic organisms are frequently interrupted by non-coding segments called introns. Before the protein can be produced, the entire length of each gene including the introns, is transcribed to produce precursor messenger RNA (pre-mRNA). An immense and dynamic molecular machine known as the spliceosome then removes the introns and splices the coding segments together to form mature mRNA, which can then be translated into proteins. A joint team from Kiyoshi Nagai’s and Sjors Scheres’ groups in the LMB’s Structural Studies Division has used electron cryo-microscopy (cryo-EM) to reveal the detailed organisation of the spliceosome.
Like other intricate molecular machines such as the ribosome, the spliceosome uses a clockwork-like mechanism to perform its vital cell function, and has many parts. The spliceosome is made from subunits called small nuclear ribonucleoprotein particles (snRNPs), the largest of which is the U4/U6.U5 tri-snRNP subunit. This large complex contains 3 RNA components and more than 30 proteins and associates with smaller snRNPs and the pre-mRNA, to form the spliceosome.
To understand the structure of this huge complex, and therefore deduce how it might function, Thi Hoang Duong (Kelly) Nguyen purified minute quantities of U4/U6.U5 tri-snRNP particles from hundreds of litres of yeast culture and, with the help of Xiaochen Bai, used electron microscopy to obtain a cryo-EM map. Kelly Nguyen and Wojtek Galej then fitted all the known crystal structures and homology models of the proteins together with the 3 RNA components into this map to show how the molecules are arranged in the complex.
This intricate structure shows the active site of the spliceosome and reveals how the spliceosome could be activated. A number of diseases including degenerative eye disease, retinitis pigmentosa, and some cancers are linked to mutations in splicing factors, which lead to errors in pre-mRNA splicing. This work helps towards understanding the molecular pathology of these diseases and could lead to possible treatments in the future.
This work was funded by the MRC, a Herchel Smith Research Studentship and a European Union Marie Curie Fellowship.