 The atomic structure of the 170 kDa membrane protein gamma-secretase, a membrane protein complex that has an important role in Alzheimer’s disease, has been solved using single-particle electron cryo-microscopy (cryo-EM) by Sjors Scheres’ group in the LMB’s Structural Studies Division. This demonstrates for the first time that high-resolution reconstruction of such small molecules can be achieved using cryo-EM.
The atomic structure of the 170 kDa membrane protein gamma-secretase, a membrane protein complex that has an important role in Alzheimer’s disease, has been solved using single-particle electron cryo-microscopy (cryo-EM) by Sjors Scheres’ group in the LMB’s Structural Studies Division. This demonstrates for the first time that high-resolution reconstruction of such small molecules can be achieved using cryo-EM.
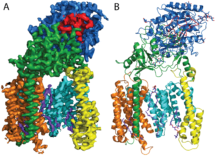
Previously, Sjors’ group had resolved the structure of human gamma-secretase to 4.5 Angstroms. By further developing cryo-EM image processing methods the research team have now resolved the structure to 3.4 Angstroms (atomic resolution). With a molecular weight of 170 kDa, gamma-secretase is the smallest single-particle structure solved to atomic resolution thus far, a milestone for cryo-EM structure determination. The atomic structure provides first-time insights into how gamma-secretase’s four components (presenilin, nicastrin, Aph-1 and Pen-2) come together to form an intra-membrane protease complex, which has a remarkably flexible active site.
Scientists have tried for decades to get a crystal structure of this complex, without success. The development of better electron detectors, driven by the LMB’s Richard Henderson, that provide less noisy images and advances in image processing software by Sjors, Yifan Cheng (UCSF) and Niko Grigorieff (HHMI Janelia Farm, LMB alumnus) have provided the technological advances needed to solve the atomic structure of such a small and non-symmetrical molecule. Xiao-chen Bai, Sjors and coworkers applied purified samples of wild type gamma-secretase, provided by Yigong Shi from Tsinghua University in Beijing, to cryo-EM grids, imaged them in the LMB’s Titan Krios microscope and solved the structure using improved image-processing tools (movie-processing algorithms, regularized likelihood optimization using the program RELION developed by Sjors, and better 3D-classification). These tools correct for movements of the proteins that are caused by the incoming electron beam, automatically determine the optimal weights for alignment and reconstruction of particles, and enable the ‘good particles’ (i.e. images of proteins that yield high-resolution reconstructions) to be separated from the ‘bad particles’ (i.e. junk, damaged or otherwise unsuitable particles). The latter was important for gamma-secretase and allowed the group to find 160 thousand ‘good particles’ from an initial data set of almost 2 million.
The possibility of solving structures of relatively small and unmodified membrane proteins is exciting: many membrane proteins are important drug targets. Gamma-secretase itself is an intra-membrane protease that cuts the amyloid precursor protein (APP), thus creating amyloid beta peptides. These amyloid beta peptides can aggregate to form plaques in the brains of patients, a hallmark of Alzheimer’s disease, particularly when gamma-secretase produces peptides that are too long. Solving the atomic structure of gamma-secretase will enable researchers to better understand how the enzyme works and can be used for structure-based drug design, which may ultimately lead to advances in the treatment for Alzheimer’s patients.
This research was funded by the MRC, an EU Marie Curie Fellowship, the Ministry of Science and Technology and the National Natural Science Foundation of China.
Further References:
Paper in Nature
Sjors’s Group Page
Yigong Shi’s Group Page
Insight on Research, July 2014: ‘Revealing the secrets of human gamma-secretase by cryo-EM’