The first structure of the FA core complex provides new insights into its function in DNA crosslink repair and why disease-causing mutations are only seen in some parts of the complex
Our DNA contains all of the information required to tell a cell what it needs to do, but it is constantly being damaged. This damage can cause severe problems, making repair processes hugely important. One common type of DNA damage, known as crosslinking, involves links forming inappropriately between two nucleotide letters. Although the specific repair pathway that fixes DNA crosslinks, and the complex at the heart of it, have been known about for decades, a full mechanistic understanding has been missing. Lori Passmore’s group, in the LMB’s Structural Studies Division, has now revealed the structure of the complex at the heart of this repair pathway for the first time.
The fundamental importance of DNA repair pathways is evident by the occurrence of diseases when these pathways are defective, including cancer. Fanconi anaemia (FA) is a severe developmental disease resulting from defects in the FA DNA repair pathway that is responsible for correcting DNA crosslinks. The FA core complex at the centre of the pathway is a multi-protein complex containing at least eight different protein subunits. It is known to create a signal at the site of DNA damage to recruit repair enzymes, but the precise mechanism has been unclear without a structure.
What is a DNA crosslink?
Linkages can form between two nucleotides of DNA within the same strand or between opposite strands of the double-stranded helix, as a result of exposure to exogenous chemicals, such as chemotherapeutic drugs and alcohol, or endogenous by-products of normal cellular metabolism.
These linkages can distort the shape of the DNA helix or prevent two strands from unwinding during DNA replication or transcription. If unrepaired, crosslinks can therefore block DNA from being replicated or transcribed to produce mRNA and then protein. Detection of unrepaired DNA crosslinks can trigger cell death. If crosslinks are found by the FA core complex, DNA repair enzymes are recruited and cell death can be avoided.
A decade in the making
This project started approximately 10 years ago when Lori first began to collaborate with KJ Patel in the LMB’s PNAC Division. KJ was working on the genetics and cell biology of FA and Lori had earlier studied a related complex (APC/C) during her PhD, so they started to work together to try to understand the mechanisms of the FA core complex. This led to the first purification of the intact natural complex, providing new insight into how it works. However, the quantity and quality of the purified protein was insufficient for high resolution structure determination.
Eeson Rajendra, a former postdoctoral researcher in Lori’s group, then developed a method for producing and purifying large quantities of the FA core complex in insect cells. Along with two other postdoctoral scientists from Lori’s group, Shabih Shakeel and Pablo Alcón, as well as through collaborations with Carol Robinson at the University of Oxford, Juri Rappsilber at Technische Universität Berlin, and the LMB’s Biological Mass Spectrometry and Proteomics Facility, the team have now determined the first structure of the FA core complex through a combination of electron cryo-microscopy and mass spectrometry.
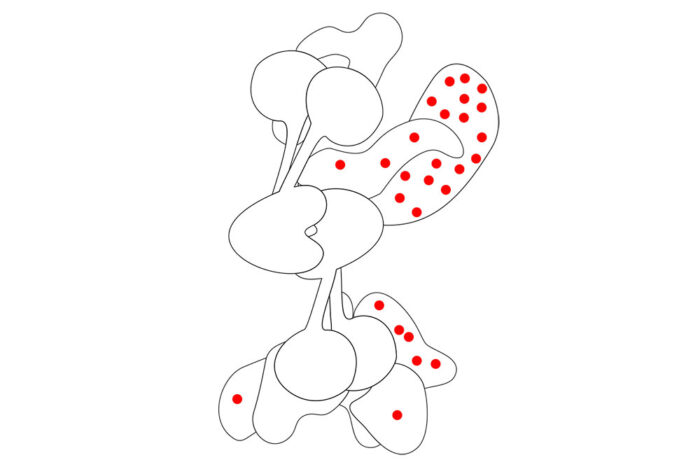
What can the structure tell us?
On close inspection, the team found that disease-causing mutations are only very rarely found at the centre of the complex. They suggest that this is due to an absolute requirement for the central part to hold the entire complex together and allow it to be functional. It might be expected that the areas where disease-causing mutations are found are the most important parts of the protein, but this is an error of survivorship bias.
A classic example of survivorship bias is said to have occurred when military concentrated on damage found on WW2 planes returning home after being subjected to enemy fire. They realised that, rather than reinforcing the areas that sustained damage, they should reinforce regions where bullet holes were not seen, as the planes that received hits in those areas simply did not return. In the same way, mutations in the core of the FA complex are not tolerated, whereas mutations elsewhere in the complex can occur and lead to disease.
As well as providing insight into the disease caused by defects in the FA pathway, understanding the mechanism by which the FA core complex recruits repair enzymes to damaged DNA is important due to the fundamental role of DNA repair. This new model will provide a basis for future investigations into the mechanisms of crosslink detection and recruitment of repair enzymes.
The work was funded by the MRC, ERC, Wellcome Trust, and BBSRC.
Further references
Structure of the Fanconi anaemia monoubiquitin ligase complex. Shakeel, S., Rajendra, E., Alcón, P., O’Reilly, F., Chorev, DS., Maslen, S., Degliesposti, G., Russo, CJ., He, S., Hill, CH., Skehel, M., Scheres, SHW., Patel, KJ., Rappsilber, J., Robinson, CV., Passmore, LA. Nature [Epub ahead of print]
Lori’s group page
Carol Robinson’s group page
Juri Rappsilber’s group page
Previous Insight on Research: Insights into how the Fanconi Anaemia core complex activates DNA repair