Time-lapse and quantitative study of developing salivary glands in Drosophila reveals key factors behind behavioural transition of epithelial cells that drives tube formation
The development of complex organs starts from simple tissue precursors – primordia – which are often flat epithelial sheets of cells. Though there has been a great deal of research to better understand how the fate of a primordium can be specified at the transcriptional level, the question of how these transcriptional cues are translated into dynamic, physical changes that sculpt the organ has remained unclear. Now, Katja Röper’s group have collaborated with Guy Blanchard from the Department of Physiology, Development and Neuroscience, University of Cambridge to use salivary glands in Drosophila embryos as a model to study the process of tubular organ formation. Their findings show that the pattern of activity of two transcription factors, Huckebein (Hkb) and Forkhead (Fkh), prefigures where a downstream key signalling molecule activates the cell shape change which leads to tube invagination.
This research builds upon recent research from Katja’ group, published in 2018, which revealed the patterning of key cell behaviours driving the earliest stages of tube budding. Interestingly, although tubular organs are symmetrical in nature, the results of the study showed that the point of invagination from the circular, epithelial primordium of embryonic salivary glands is in an asymmetric position, rather than central to the placode. Now, Katja’s group have revealed how this asymmetry is established and maintained during the formation of salivary gland tubes.
Yara Sanchez-Corrales, a postdoc in Katja’s lab, tagged endogenous proteins to label cell shapes in Drosophila embryos. She then took time-lapse movies of the developing salivary glands. These movies were then analysed using computational tools and quantitative approaches to identify how cells – and the tissue overall – change throughout organ formation. This revealed that apical constriction of cells was strongest closest to the invagination pit (the point of folding into a tubular structure) at any one time. In fact, the movies allowed the group to witness how cells switch their behaviour from mostly intercalating to apically constricting as they approached the region near the pit.
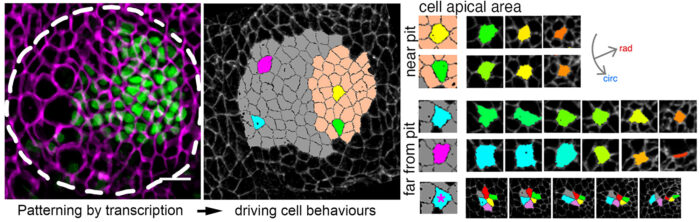
The impetus behind this transition was discovered when Yara fluorescently tagged the transcription factors, HkB and Fkh. The team observed that the expression levels of these proteins are greatest where the initial constriction occurs. Both operate upstream of the expression of the GPCR ligand Fog, which in turn activates apical-medial myosin.
To confirm the importance of these transcription factors, the team repeated the experiments in Drosophila embryos which were modified without the correct pre-patterning of Hkb or Fkh, or with an overexpression of Fog. In these instances, this led to a non-symmetrical final organ shape with widened cavities and sack-like glands, thereby reinforcing their earlier observations that these are the driving factors behind symmetrical morphogenesis of tubular organs.
The study of tube formation holds huge importance as most of our internal organs are tubular in nature, such as the lungs, kidneys, our intestines, the pancreas and mammary glands. Research to better understand the process and the control factors of organ formation thus holds the potential to eventually shed light on how congenital malformations occur and how failures in tubular tissue homeostasis arise, paving the way for treatments to be developed.
This work was funded by UKRI MRC and the Wellcome Trust
Further references
Correct regionalisation of a tissue primordium is essential for coordinated morphogenesis. Sanchez-Corrales, YE., Blanchard, G., Röper, K., eLife 10:e72369 (2021)
Katja Röper’s group
Guy Blanchard’s group (Department of Physiology, Development and Neuroscience, University of Cambridge)
Previous Insights on Research articles
Discovery of a key piece of the puzzle of tubular organ formation
How flat sheets of cells become tubular organs: observing cellular dynamics from 2D from 3D