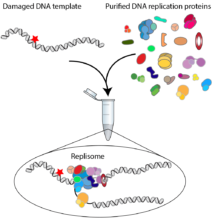
Work by Joe Yeeles’ group in the LMB’s PNAC Division has for the first time revealed the earliest responses when the eukaryotic DNA replication machinery, the replisome, collides with DNA damage.
Every time a cell divides, its DNA must be replicated so that each daughter cell inherits a complete copy of the genome. Since DNA carries the instructions needed for cells to function, DNA replication must be extremely accurate to prevent the inheritance of mutations which can lead to defects in cellular function. However, the DNA replication machinery frequently encounters obstacles that impede its progression, including DNA damage in the parental DNA strand being copied. Dealing with these obstacles is critical to ensure complete and accurate genome duplication, but the earliest events that occur after the replication machinery collides with damaged DNA were, until now, very poorly understood in eukaryotic organisms.
Martin Taylor, from Joe’s group, recreated DNA replication in a test tube, using purified yeast replication proteins and yeast DNA templates. He engineered DNA damage into the parental DNA strand and observed what happened when the replication protein machinery tried to replicate this DNA. Martin and Joe found that the response of the replisome absolutely depended on whether the damage was present in the continuously-replicated “leading strand” template or the discontinuously-replicated “lagging strand” template.
If the damage was replicated by the lagging strand, the replisome simply jumped past the damage without delay leaving a very short un-replicated gap behind. This is because during discontinuous lagging strand replication under normal (undamaged) conditions, the “primase” enzyme DNA polymerase alpha-primase repeatedly initiates synthesis of short fragments of DNA which get joined up later, and so the damage only inhibits one of these short fragments.
In contrast, if the damage was replicated by the leading strand, collisions with damage caused very severe delays in replisome progression and the accumulation of abnormal DNA structures. Martin and Joe found for the first time that the yeast replisome can re-establish a new leading strand beyond the damage without the assistance of accessory factors, via the activity of the primase on the leading strand. Surprisingly, this activity was extremely inefficient even after multiple sites of DNA damage in the same molecule – in complete contrast to the bacterial replisome, which was shown to perform such reactions efficiently in earlier studies led by Joe during his first postdoctoral position in Ken Marians group in the US. Martin and Joe then pinpointed down the source of this inefficiency to the initial establishment of the new leading strand by the primase. They also found that under certain conditions, which partly mimic the situation where cells are exposed to high doses of DNA damage during DNA replication, the restart reaction can become more efficient.
Taken together the results show that the action of the primase enzyme is very different on the leading and lagging strands, and this has significant implications for the control of damage bypass in different contexts. The development of the experimental system also provides a platform to ask how other proteins known to control the response of replisomes to DNA damage work in vivo. By adding such proteins into the system, scientists can investigate how the proteins associate with stalled replication forks to help bypass of damage, important for accurate DNA replication and avoidance of disease.
This work was funded by the MRC and Wellcome Trust.