The entire neuropeptide signalling network of a whole animal, the nematode worm C. elegans, has been mapped for the first time
Understanding how the brain functions remains one of the biggest challenges in biology. Efforts are ongoing to build wiring diagrams of the brain, revealing the synaptic connections between neurons, but this is only one way that signals flow through the nervous system. Another key neural communication pathway is via signalling molecules – in particular via neuropeptides – that modulate synaptic communication and allow communication between neurons that are not linked by synapses.
Although these wireless networks are extensive, complex, and critical for brain function, they have never before been characterised in any animal in this detail. Now, for the first time the entire wireless neuropeptide network of a whole animal has been mapped by William Schafer’s group in the LMB’s Neurobiology Division, in collaboration with an international team including Petra Vértes’ group in Department of Psychiatry, University of Cambridge and Isabel Beets’ group in KU Leuven. Professor Beets is also an LMB Fellow in the Neurobiology Division.
Lidia Ripoll-Sánchez, a Ph.D. student in the Schafer group, pieced together the wireless network of the nematode worm, C. elegans, by integrating single-cell anatomical and gene expression datasets with biochemical analysis of receptor-ligand interactions. This enabled Lidia to determine which neuropeptides and receptors are expressed in each of the 302 neurons of C. elegans, and which neuropeptides interact with which receptor. By combining this information with the anatomy of the nematode worm she could then infer which neurons send peptidergic signals to which targets and build the whole animal neuropeptide signalling network, comprising 31,478 connections.
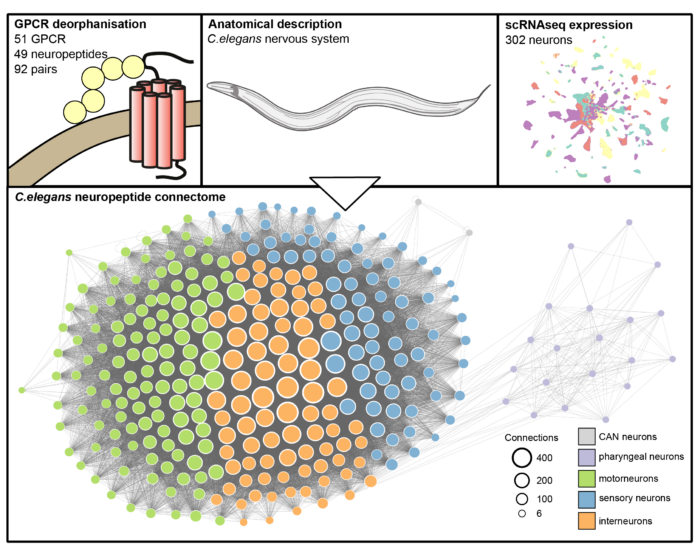
Once the neuropeptide network had been mapped, the team used graph theory to analyse its physical properties and compared it to the synaptic connectome. They identified several interesting and unexpected features: the neuropeptide network is denser, more decentralised, and the most important hubs are different from the main hubs of the synaptic network. The neuropeptide network also integrates parts of the nervous system that are isolated from the synaptic connectome.
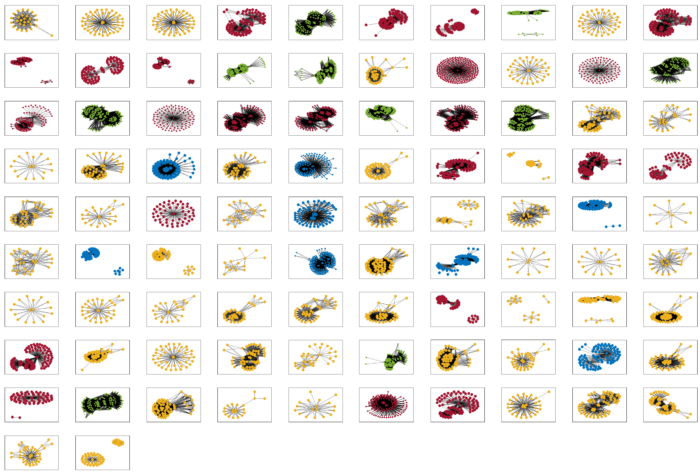

This is the first time the neuropeptide signalling network for a whole nervous system has been analysed in this way. Although the C. elegans nervous system is anatomically small, at the molecular level its neuropeptide systems are highly complex and show significant homology to other animals. Since the synaptic connectome of the nematode worm shows many features that are conserved in bigger brains, the team expect that the C. elegans neuropeptide connectome will also serve as a prototype to understand wireless signalling in larger nervous systems.
Neuropeptide systems play important roles in behavioural states including sleep, feeding, reproduction and learning. In humans, neuropeptide receptors are the target of drug development for new neuropsychiatric treatments for conditions including insomnia, chemotherapy-induced nausea, schizophrenia and depression. But the way these drugs act in the brain at the network level is not well understood. The structure of neuropeptide networks suggests that they may process information in a different way to synaptic networks. Understanding how this works may not only help us understand how drugs work but also how our emotions and mental states are controlled.
This work was funded by UKRI MRC, NIH, the Wellcome Trust, MQ: Transforming Mental Health, the ERC, HHMI, KU Leuven and the Simons Foundation.
Further references
The neuropeptidergic connectome of C. elegans Ripoll-Sánchez, L., Watteyne, J., Sun, HS., Fernandez, R., Taylor, SR., Weinreb, A., Bentley, BL., Hammarlund, M., Miller, DM., Hobert, O., Beets, I., Vértes, PE., Schafer, WR. Neuron
William’s group page
Petra Vértes’ page
Isabel Beets’ page
UKRI news release
Previous Insight on Research articles
Ankyrins: a missing molecular link in mechanical senses
How neuropeptide signalling controls sensitisation in response to touch in C. elegans
Network control principles predict neuron function in the C. elegans connectome