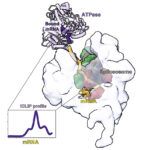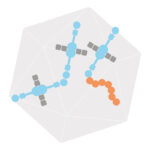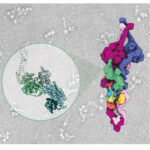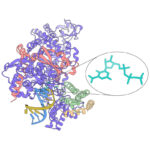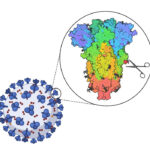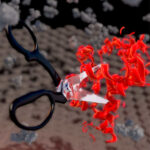
A powerful strategy to study protein function has been to deplete a protein of interest from the cell and then study the consequences. Leo James’ group has exploited new understanding of TRIM21’s mechanism of activation to develop new tools for targeted protein degradation.