Biophysics
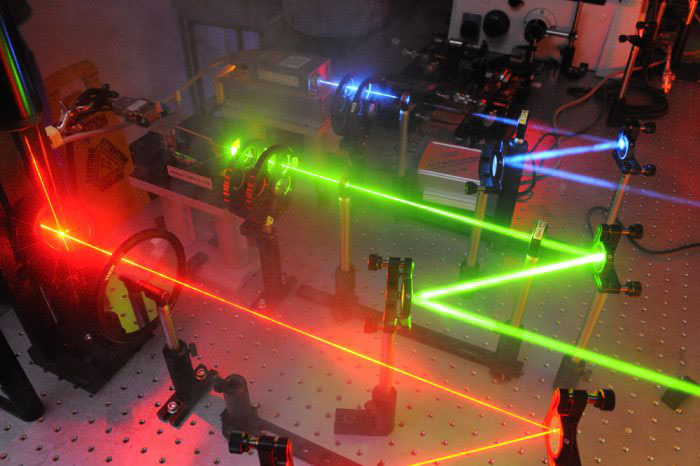

The biophysics facility brings established theories and methods from the physical sciences and applies these to open questions in structural biology. Biophysical techniques contribute to all areas of research in the LMB from simple expression and purification issues through to verifying high-resolution structural information and probing interactions and biology from single molecules through to whole cells.
Researchers can access a wide range of biophysical techniques that probe the hydrodynamic, spectroscopic, thermodynamic and physical properties of biological macromolecules and their interactions. Techniques covered include sensor based platforms such as Biacore, Octet and SwitchSense, titration and scanning calorimetry, analytical centrifugation, circular and linear dichroism, fluorescence and scanning fluorimetry, static and dynamic light scattering and microscale thermophoresis. In addition, there are also a number of ‘in-house’ instruments for performing the latest cutting edge techniques in rapid reaction relaxation kinetics and single molecule spectroscopies. Many of these methods are central to the success of research programmes in the LMB.
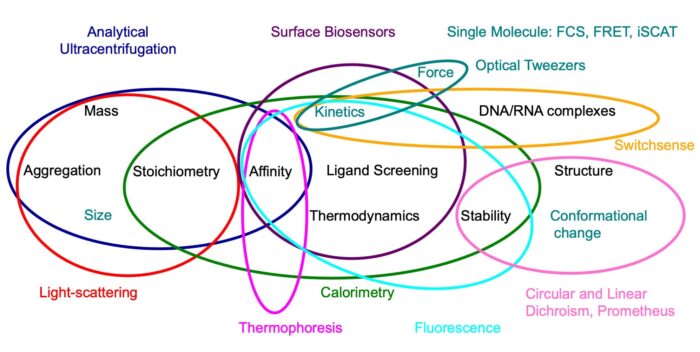
These world-class facilities have an expert team of scientists headed by Stephen McLaughlin, supported by Chris Johnson and Chris Batters, who bring many years of experience with these techniques in numerous applications.. To help researchers make their own routine measurements, the biophysics team provides an annual lecture series on methods and instrumentation within the LMB as well as offering detailed one-to-one training as required. In addition, these experts can discuss the design of proposed experiments and advise on the interpretation and fitting of results. The team also engages in fully collaborative projects where there is a requirement for specialist skills in the design and execution of the work and where analysis and fitting of data are complex.
In collaboration with AstraZeneca and Imperial College, London, the facility organises an annual one-day symposium on Next Generation Biophysics, that brings together scientists in both academia and industry for stimulating talks examining the application of cutting edge biophysical techniques in complex biological settings.
Selected Papers
Yatskevich, S., Muir, K.W., Bellini, D., Zhang, Z., Yang, J., Tischer, T., Predin, M., Dendooven, T., McLaughlin, S.H., Barford, D. (2022)
Structure of the human inner kinetochore bound to a centromeric CENP-A nucleosome.
Science: 376(6595):844-852. doi: 10.1126/science.abn3810
Johnson, C.M. (2021)
Isothermal Titration Calorimetry in Protein-Ligand Interactions: Methods and Applications
Methods in Molecular Biology (2263) 3rd ed. (Daviter, T., Johnson, C.M., McLaughlin, S.H., and Williams, M.A., eds), Humana Press (USA).
Fischer, E.S., Yu, C.W.H., Bellini, D., McLaughlin, S.H., Orr, C.M., Wagner, A., Freund, S.M.V., Barford D. (2021)
Molecular mechanism of Mad1 kinetochore targeting by phosphorylated Bub.
EMBO Rep 22(7): e52242. doi: 10.15252/embr.202052242.
Ohashi, Y., Tremel, S., Masson, G.R., McGinney, L., Boulanger, J., Rostislavleva, K., Johnson, C.M., Niewczas, I., Clark, J., Williams, R.L. (2020)
Membrane characteristics tune activities of endosomal and autophagic human VPS34 complexes,
Elife. 9:e58281. doi: 10.7554/eLife.58281.
Kiss, L., Zeng, J., Dickson, C.F., Mallery, D.l., Yang, C.-J., McLaughlin, S.H., Boland, A., Neuhaus, A., James, L.C. (2019)
A tri-ionic anchor mechanism drives Ube2N-specific recruitment and K63-chain ubiquitination in TRIM ligases.
Nature Commun: 10: 4502 doi: 10.1038/s41467-019-12388-y.
Inglis, A. J., Masson, G. R., Shao, S., Perisic, O., McLaughlin, S. H., Hegde, R. S. and Williams, R. L. (2019)
Activation of GCN2 by the ribosomal P-stalk.
Proc Natl Acad Sci U S A: 116(11):4946-4954. doi: 10.1073/pnas.1813352116. Epub 2019 Feb 25.
Liu Y., Gupta G.D., Barnabas D.D., Agircan F.G., Mehmood S., Wu D., Coyaud E., Johnson C.M., McLaughlin S.H., Andreeva A., Freund S.M.V., Robinson C.V., Cheung S.W.T., Raught B., Pelletier L., van Breugel M. (2018)
Direct binding of CEP85 to STIL ensures robust PLK4 activation and efficient centriole assembly.
Nat Commun. 9(1):1731
Izoré T., Kureisaite-Ciziene D., McLaughlin S.H., Löwe J. (2016)
Crenactin forms actin-like double helical filaments regulated by arcadin-2.
J. Elife. 5.
Gammons M.V., Renko M., Johnson C.M., Rutherford T.J., Bienz M. (2016)
Wnt Signalosome Assembly by DEP Domain Swapping of Dishevelled.
Mol Cell. 64(1):92-104.
Chang L., Zhang Z., Yang J., McLaughlin S.H., Barford D. (2015)
Atomic structure of the APC/C and its mechanism of protein ubiquitination.
Nature. 522(7557):450-454.
Johnson C.M. (2013)
Differential scanning calorimetry as a tool for protein folding and stability.
Arch Biochem Biophys. 531(1-2):100-9.