

 Hugh Pelham obtained his degree and PhD in Biochemistry from Cambridge University, and after postdoctoral studies with Donald Brown in Baltimore joined the LMB as a group leader in 1981. He was Head of the Cell Biology Division (initially Joint Head) from 1992 to 2006, Deputy Director from 1996 to 2006, and Director from 2006 to 2018. As Deputy Director and Director he was deeply involved in the design and delivery of the new building to which LMB moved in 2013.
Hugh Pelham obtained his degree and PhD in Biochemistry from Cambridge University, and after postdoctoral studies with Donald Brown in Baltimore joined the LMB as a group leader in 1981. He was Head of the Cell Biology Division (initially Joint Head) from 1992 to 2006, Deputy Director from 1996 to 2006, and Director from 2006 to 2018. As Deputy Director and Director he was deeply involved in the design and delivery of the new building to which LMB moved in 2013.
As a group leader, Hugh worked initially on the transcriptional control of heat shock genes. Turning to the role of the heat shock proteins themselves, he proposed a universal function in protein folding, with properties now encapsulated in the term “molecular chaperone”. The identification of chaperones resident in the Endoplasmic Reticulum led to the discovery of the KDEL retention system, and the KDEL receptor. This was the starting point for wider studies, mostly in yeast, of protein sorting and trafficking. His group characterised many of the SNARE proteins that catalyse vesicle fusion, including the sorting signals that ensure their correct location, and the transport pathways and coat proteins that mediate this. More recently he studied the role of ubiquitination in membrane protein sorting and quality control, and the role of adaptor proteins in controlling the activity of NEDD4 family ubiquitin ligases, and guiding them to their substrates.
 Hugh is an honorary Fellow of Christ’s College, and an emeritus honorary Professor of Molecular Biology at the University of Cambridge. He was elected a Member of the European Molecular Biology Organisation in 1985, a Fellow of the Royal Society in 1988, and was a founder Fellow of the Academy of Medical Sciences in 1998. He has received a number of scientific awards, including the EMBO gold medal, the Royal Society Croonian Lecture and medal, the Louis Jeantet Prize for Medicine and the King Faisal International Prize for Science, and was awarded a Knighthood for services to science in 2011.
Hugh is an honorary Fellow of Christ’s College, and an emeritus honorary Professor of Molecular Biology at the University of Cambridge. He was elected a Member of the European Molecular Biology Organisation in 1985, a Fellow of the Royal Society in 1988, and was a founder Fellow of the Academy of Medical Sciences in 1998. He has received a number of scientific awards, including the EMBO gold medal, the Royal Society Croonian Lecture and medal, the Louis Jeantet Prize for Medicine and the King Faisal International Prize for Science, and was awarded a Knighthood for services to science in 2011.
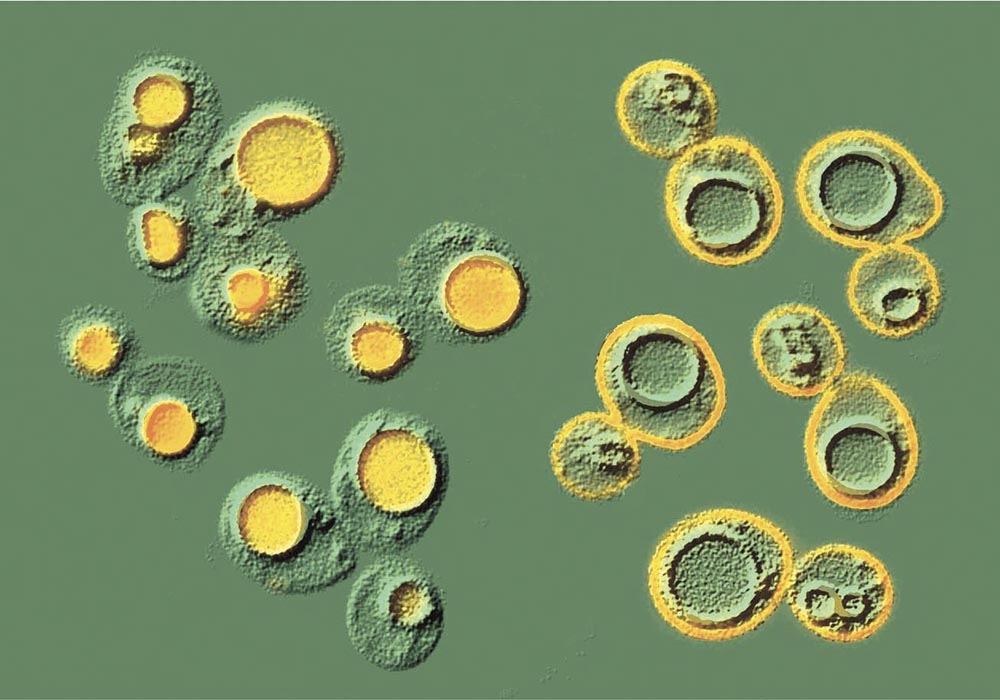
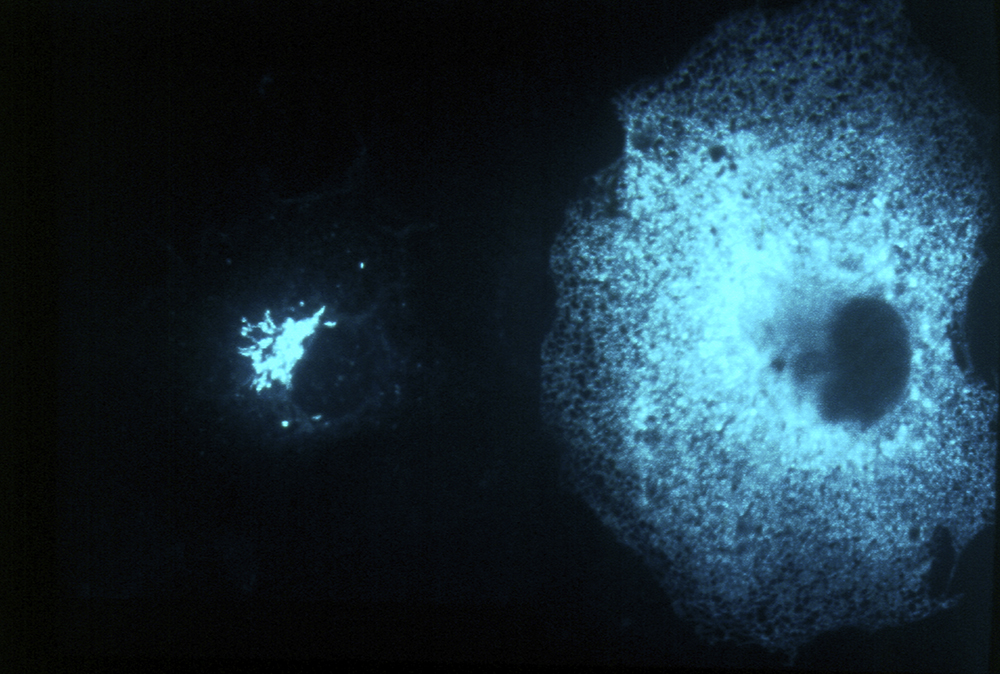
Eukaryotic cells are subdivided into membrane-bounded organelles, each with a characteristic set of membrane proteins. These proteins must be targeted to their correct locations, a process which involves their segregation into small vesicles which bud from one compartment and fuse with another.
Equally important for the cell is the ability to remove unwanted or damaged proteins. Misfolded proteins are frequently degraded in the endoplasmic reticulum, but once proteins leave the ER the main route to destruction is via the multivesicular body pathway to lysosomes or vacuoles. This is the fate of proteins whose degradation is regulated – such as activated signalling receptors, or excess transporters and channels. These proteins are marked for degradation by addition of the small protein ubiquitin to their cytoplasmic domains, the attached tag acting as a sorting signal for the multivesicular body pathway.
We have studied the mechanisms by which such proteins are recognised, modified and sorted, and how the activities of the ubiquitin ligases are regulated.
Selected Papers
- Mund T., Pelham H. R. (2018)
Substrate clustering potently regulates the activity of WW-HECT domain-containing ubiquitin ligases
Journal of Biological Chemistry 293(14): 5200-5209. - Mund T., Graeb M., Mieszczanek J., Gammons M., Pelham H. R., Bienz M. (2015)
Disinhibition of the HECT E3 ubiquitin ligase WWP2 by polymerized Dishevelled
Open Biology 5(12): 150185. - Mund, T., Lewis, M.J., Maslen, S. and Pelham, H.R.B (2014)
Peptide and small molecule inhibitors of HECT-type ubiquitin ligases.
Proc Natl Acad Sci USA 111: 16736-16741. - Mund, T. and Pelham, H.R.B. (2010)
Regulation of PTEN/Akt and MAP kinase signalling pathways by the ubiquitin ligase activators Nfip1 and Ndfip2.
Proc Natl Acad Sci USA 107: 11429-11434.