

Kelly Nguyen
Molecular mechanism of telomere maintenance and the roles of telomeres in human diseases
knguyen@mrc-lmb.cam.ac.ukPersonal group site
The ends of linear eukaryotic chromosomes are capped telomeres, an array of long telomeric repeat tracts ([TTAGGG]nin human) bound by telomeric protein factors. Telomeres distinguish natural chromosomal ends from DNA breaks and thereby protect chromosomes against degradation and interchromosomal fusion. Each cell division results in the loss of a small portion of telomeres as a consequence of incomplete genome replication. Critically short telomeres lose their end-protection capability, resulting in genome instability, proliferative senescence or cell death. Therefore, telomere length is often regarded as a “cellular aging clock” and telomere maintenance is essential for genome integrity. Telomere dysfunction has been linked to cancers and premature aging syndromes such as dyskeratosis congenita, aplastic anemia, and pulmonary fibrosis in human patients.
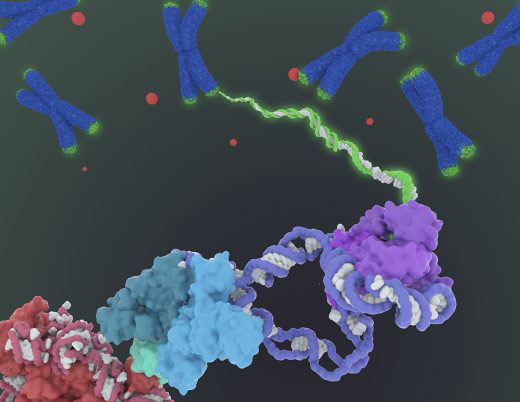
We developed biochemical methods to reconstitute human telomerase holoenzyme, a large ribonucleoprotein that compensates for telomere shortening by synthesizing the 3’ telomeric DNA repeats, and determined its cryo-EM structure at medium resolution. The structure settled the longstanding question regarding its composition and gave unprecedented insight into its assembly.
We are studying telomerase regulation at telomeres and other processes involved in telomere maintenance beyond telomerase. We employ an integrated structural biology approach with a focus on biochemistry, cryo-electron microscopy and in vivo studies in mammalian cells. Ultimately, our aim is to uncover how mutations in human patients result in telomere dysfunction, which will be essential for telomere-based therapeutics.
Selected Papers
- Ghanim, G.E., Hu, H., Boulanger, J., Nguyen, T.H.D. (2025)
Structural mechanism of LINE-1 target-primed reverse transcription.
Science 388(6745): eads8412 - Hu, H; van Roon, AMM; Ghanim, GE; Ahsan, B; Oluwole, AO; Peak-Chew, S-Y; Robinson, CV; Nguyen, THD (2023)
Structural basis of telomeric nucleosome recognition by shelterin factor TRF1
Sci Adv 9(34): eadi4148. - Sekne, Z., Ghanim, GE., van Roon, AMM., Nguyen, THD. (2022)
Structural basis of human telomerase recruitment by TPP1-POT1
Science 375(6585): 1173-1176. - Ghanim, G.E., Fountain, A.J., van Roon, A.M., Rangan, R., Das, R., Collins, K. and Nguyen, T.H.D. (2021)
Structure of human telomerase holoenzyme with bound telomeric DNA
Nature 593(7859): 449-453 - Nguyen THD, Tam J, Wu RA, Greber BG, Toso D, Nogales E, Collins K. (2018)
Cryo-EM structure of substrate-bound human telomerase holoenzyme.
Nature 557: 190-195. - Nguyen THD, Galej WP, Bai X.-C., Oubridge C, Newman AJ, Scheres SHW, Nagai K. (2016)
CryoEM structure of the yeast U4/U6.U5 tri-snRNP at 3.7 Å resolution.
Nature 530: 298-302. - Nguyen THD, Galej WP, Bai X.-C, Savva CG, Newman AJ, Scheres SHW, Nagai K. (2015)
The architecture of the spliceosomal U4/U6.U5 tri-snRNP.
Nature 523: 47-52.
Group Members
- Naura Antariksa
- Sebastian Balch
- Nirupa Desai
- Elsa Franco Echevarria
- Inga Hochheiser
- Hongmiao Hu
- Simonas Melaika
- Martin Saurer
- Elizabeth Wang
- Helen Yan